Difference between revisions of "MiPNet21.06 SUIT RP"
| Line 111: | Line 111: | ||
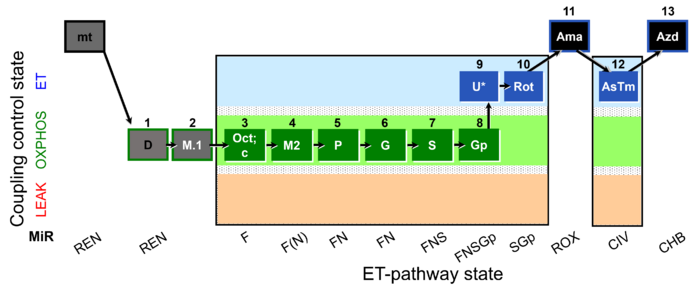
== SUIT reference protocol RP2 == | == SUIT reference protocol RP2 == | ||
[[File:1D;1M.1;2Oct;2c;2M2;3P;4G;5S;6Gp;7U;8Rot;9Ama;10Tm;11Azd.jpg|right|700px|link=SUIT-RP2 |SUIT-RP2]]<br /> | |||
****:» [[1D;2OctM;3P;4G;5S;6Gp;7U;8Rot-]] | ****:» [[1D;2OctM;3P;4G;5S;6Gp;7U;8Rot-]] | ||
'''SUIT states:''' 1[[ROX]] 2[[OctM]](P) 3[[OctM]](c) 4[[OctPM]] 5[[OctPGM]] 6[[OctPGMS]] 7[[OctPGMSGp]] 8[[E]] 9[[SGp]] 10[[ROX]] 11[[CIV]] 12[[ROX]] | '''SUIT states:''' 1[[ROX]] 2[[OctM]](P) 3[[OctM]](c) 4[[OctPM]] 5[[OctPGM]] 6[[OctPGMS]] 7[[OctPGMSGp]] 8[[E]] 9[[SGp]] 10[[ROX]] 11[[CIV]] 12[[ROX]] | ||
Revision as of 12:48, 16 March 2017
| SUIT reference protocol for OXPHOS analysis by high-resolution respirometry. |
» ![]()
OROBOROS (2016-08-17) Mitochondr Physiol Network
Abstract: Doerrier C, Sumbalova Z, Krumschnabel G, Hiller E, Gnaiger (2016) SUIT reference protocol for OXPHOS analysis by high-resolution respirometry. Mitochondr Physiol Network 21.06(01):1-12. » [In progress]
- » MitoPedia: SUIT reference protocol
- » Instrument: O2k, O2k-Catalogue
• O2k-Network Lab: AT_Innsbruck_OROBOROS
Labels: MiParea: Instruments;methods
Preparation: Permeabilized cells, Permeabilized tissue, Homogenate, Isolated mitochondria
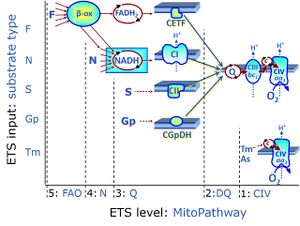
Regulation: Substrate Coupling state: LEAK, OXPHOS, ETS"ETS" is not in the list (LEAK, ROUTINE, OXPHOS, ET) of allowed values for the "Coupling states" property. Pathway: F, N, S, Gp, CIV, NS, Other combinations HRR: Oxygraph-2k, O2k-Protocol
MitoPathways, MitoFitPublication, 1PM;2D;3U;4G;5S;6Oct;7Rot;8Gp-, 1D;2OctM;3P;4G;5S;6Gp;7U;8Rot-
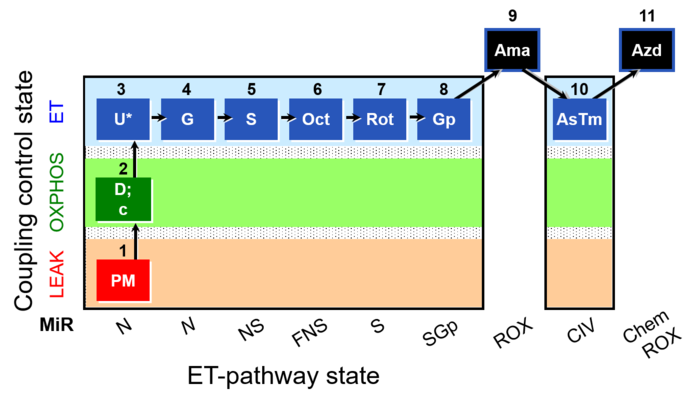
SUIT reference protocol RP1
SUIT states: 1-4PM(LPcE) 5PGM 6PGMS 7OctPGMS 8S 9SGp 10ROX 11Tm 12ROX
| Step | Respiratory state | Pathway control | Pathway to Q |
|---|---|---|---|
| 1PM | PM(L) | N | CI |
| 2D | PM(P) | N | CI |
| 3c | PM(P)c | N | CI |
| 3U | PM(E) | N | CI |
| 4G | PGM(E) | N | CI |
| 5S | PGMS(E) | NS | CI&II |
| 6Oct | OctPGMS(E) | FNS | FAO&CI&II |
| 7Rot | S(E) | S | CII |
| 8Gp | SGp(E) | SGp | CII&GpDH |
| 9Ama | ROX | ROX | |
| 10Tm | Tm(E) | CIV | CIV |
| 11Azd | ROX | ROX |
SUIT reference protocol RP2
SUIT states: 1ROX 2OctM(P) 3OctM(c) 4OctPM 5OctPGM 6OctPGMS 7OctPGMSGp 8E 9SGp 10ROX 11CIV 12ROX
| Step | Respiratory state | Pathway control | Pathway to Q |
|---|---|---|---|
| 1D | ROX | ROX | |
| (M.1) | |||
| 2Oct | Oct(P) | (F) | FAO |
| 2c | OctM(P)c | F | FAO |
| 2M2 | OctM(P) | F | FAO |
| 3P | OctPM(P) | FN | FAO&CI |
| 4G | OctPGM(P) | FN | FAO&CI |
| 5S | OctPGMS(P) | FNS | FAO&CI&II |
| 6Gp | OctPGMSGp(P) | FNSGp | FAO&CI&II&GpDH |
| 7U | OctPGMSGp(E) | FNSGp | FAO&CI&II&GpDH |
| 8Rot | SGp(E) | SGp | CII&GpDH |
| 9Ama | ROX | ROX | |
| 10Tm | Tm(E) | CIV | CIV |
| 11Azd | ROX | ROX |
DL-Drivers and DatLab-Analysis templates SUIT protocol RP1
- SUIT RP1:1PM;2D:2c;3U;4G;5S;6Oct;7Rot;8Gp;9Ama;10Tm;11Azd
DL-Drivers and DatLab-Analysis templates SUIT protocol RP2
- SUIT RP2:1D;1M.1;2Oct;2c;2M2;3P;4G;5S;6Gp;7U;8Rot;9Ama;10Tm;11Azd
Experimental details
Work in progress: pre-publication protocol
Saturating ADP concentrations
- Different concentrations of ADP are sufficient to obtain maximum flux for estimating OXPHOS capacity. The typical range for isolated mitochondria is 1 to 2.5 mM, for permeabilized cells 1 to 5 mM, for permeabilized muscle fibres 2.5 to 10 mM.
RP-T01
- RP1: Compared to RP1-T01, move Oct titration after S.
- RP2: Compared to RP2-T01, move U titration after Gp.
RP-T02
- RP2: Compared to RP2-T02, move D titration after the sample. Titration of M.1 before Oct. M2 is added after Oct.
Cytochrome c test
- In protocol RP2, the cytochrome c test might better be moved from a c-titration after pyruvate (P) to a titration after M2. Otherwise, there is a risk of underestimation of FAO in cases of a cytochrome c effect. On the other hand, at the low flux before titration of P, the sensitivity of the cytochrome c test may be low.
- ~ Gnaiger Erich 18:26, 23 January 2016 (CET)
Mark names
- Even the highly abbreviated names of respiratory states have become too long for DatLab, where mark names are restricted to 8 digits. The visibility in DatLab of long mark names is restricted. The simplest solution for short mark names is the use of a numerical sequence, with the preceeding event name added for information.
- ~ Gnaiger Erich 16:50, 23 January 2016 (CET)
Pre-publication communication
- This communication is a pre-publication, inviting critical feedback and comments. All feedback will be carefully documented and evaluated in terms of justification of co-authorship. We intend to finally publish this topic in a peer-reviewed journal as an original article, with reference to the pre-publication history including reviewer's reports. - Gnaiger Erich 03:23, 18 January 2016 (CET)
Experiments in progress
- The SUIT reference protocol is presently applied in permeabilized HEK cells, mouse heart isolated mitochondria, liver homogenate, permeabilized skeletal muscle (mouse and human), and human PBMCs and platelets. - Gnaiger Erich 18:47, 19 January 2016 (CET)
- AT Innsbruck Gnaiger E and AT Innsbruck OROBOROS - MitoFit project.
- US NC Winston-Salem Molina AJA - UPBEAT project.
Further details
- » Introduction: Gnaiger 2014 MitoPathways
- » MitoPedia: SUIT - extended MitoPedia topic, May 2016
- Table of titrations » MiPNet09.12 O2k-Titrations
- » Definition: Substrate-uncoupler-inhibitor titration
- » Context: SUIT protocol library
- » Abbreviations: MitoPedia