|
|
| Line 7: |
Line 7: |
| ::: '''[[SUIT protocol pattern]]:''' diametral/complex 1OctM;2D;3G;4S;5Rot;6Omy;7U | | ::: '''[[SUIT protocol pattern]]:''' diametral/complex 1OctM;2D;3G;4S;5Rot;6Omy;7U |
| __TOC__ | | __TOC__ |
| Communicated by [[Doerrier C]] and [[Gnaiger E]] (last update 2019-01-16) | | Communicated by [[Doerrier C]], [[Huete-Ortega M]] and [[Gnaiger E]] (last update 2019-01-30) |
| == DLP applications == | | == DLP applications == |
|
| |
|
Revision as of 10:41, 30 January 2019

- high-resolution terminology - matching measurements at high-resolution
SUIT-016
Description
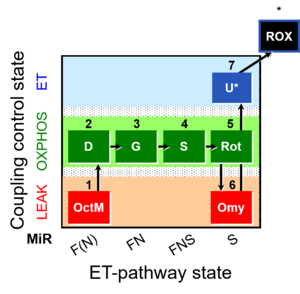
Abbreviation: FNS(Oct,GM)
Reference: A  »Versions
»Versions
- SUIT-category: FNS(Oct,GM)
- SUIT protocol pattern: diametral/complex 1OctM;2D;3G;4S;5Rot;6Omy;7U
Communicated by Doerrier C, Huete-Ortega M and Gnaiger E (last update 2019-01-30)
DLP applications
References
| | Year | Reference | Organism | Tissue;cell |
|---|
| Gnaiger 2015 Scand J Med Sci Sports | 2015 | Gnaiger E, Boushel R, Søndergaard H, Munch-Andersen T, Damsgaard R, Hagen C, Díez-Sánchez C, Ara I, Wright-Paradis C, Schrauwen P, Hesselink M, Calbet JAL, Christiansen M, Helge JW, Saltin B (2015) Mitochondrial coupling and capacity of oxidative phosphorylation in skeletal muscle of Inuit and caucasians in the arctic winter. https://doi.org/10.1111/sms.12612 | Human | Skeletal muscle |
Steps and respiratory states
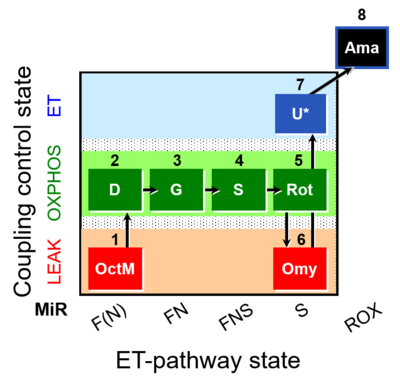
| Step
|
State
|
Pathway
|
Q-junction
|
Comment - Events (E) and Marks (M)
|
| 1OctM
|
OctML(n)
|
F(N)
|
CETF
|
1OctM
- Respiratory stimulation of the FAO-pathway, F, by fatty acid, FA, in the presence of malate, M. Malate is a type N substrate (N), required for the F-pathway. The FA concentration has to be optimized to saturate the F-pathway, without inhibiting or uncoupling respiration.
- Low concentration of malate, typically 0.1 mM, does not saturate the N-pathway; but saturates the F-pathway.
- Non-phosphorylating resting state (LEAK state); LEAK respiration L(n) in the absence of ADP, ATP, AMP (no adenylates).
|
| 2D
|
OctMP
|
F(N)
|
CETF
|
1OctM;2D
- Respiratory stimulation of the FAO-pathway, F, by fatty acid, FA, in the presence of malate, M. Malate is a type N substrate (N), required for the F-pathway. The FA concentration has to be optimized to saturate the F-pathway, without inhibiting or uncoupling respiration.
- Low concentration of malate, typically 0.1 mM, does not saturate the N-pathway; but saturates the F-pathway.
- OXPHOS capacity P (with saturating [ADP]), active OXPHOS state.
|
| 3G
|
OctGMP
|
FN
|
CETF&I
|
1OctM;2D;3G
- Respiratory stimulation of the FAO-pathway, F, by fatty acid, FA, in the presence of malate, M. Malate is a type N substrate (N), required for the F-pathway. The FA concentration has to be optimized to saturate the F-pathway, without inhibiting or uncoupling respiration. & NADH-linked substrates (type N-pathway to Q).
- Respiratory stimulation by simultaneous action of the F-pathway and N-pathway with convergent electron flow in the FN-pathway for evaluation of an additive or inhibitory effect of F.
- OXPHOS capacity P (with saturating [ADP]), active OXPHOS state.
|
| 4S
|
OctGMSP
|
FNS
|
CETF&CI&II
|
1OctM;2D;3G;4S
- Respiratory stimulation of the FAO-pathway, F, by fatty acid, FA, in the presence of malate, M. Malate is a type N substrate (N), required for the F-pathway. The FA concentration has to be optimized to saturate the F-pathway, without inhibiting or uncoupling respiration. & NADH-linked substrates (type N-pathway to Q). & Succinate, S ( type S-pathway to Q).
- Respiratory stimulation by simultaneous action of the F-pathway, N-pathway, and S-pathway, with convergent electron flow in the FNS-pathway for reconstitution of TCA cycle function and additive or inhibitory effect of F.
- OXPHOS capacity P (with saturating [ADP]), active OXPHOS state.
|
| 5Rot
|
SP
|
S
|
CII
|
1OctM;2D;3G;4S;5Rot
|
| 6Omy
|
SL(Omy)
|
S
|
CII
|
1OctM;2D;3G;4S;5Rot;6Omy
- Succinate, S ( type S-pathway to Q).
- Non-phosphorylating resting state (LEAK state); LEAK-respiration, L(Omy), after blocking the ATP synthase with oligomycin.
|
| 7U
|
SE
|
S
|
CII
|
1OctM;2D;3G;4S;5Rot;6Omy;7U
|
| 8Ama
|
ROX
|
|
|
1OctM;2D;3G;4S;5Rot;6Omy;7U;8Ama
- Rox is the residual oxygen consumption in the ROX state, due to oxidative side reactions, estimated after addition of antimycin A (inhibitor of CIII). Rox is subtracted from oxygen flux as a baseline for all respiratory states, to obtain mitochondrial respiration (mt).
|
| Step
|
Respiratory state
|
Pathway control
|
ET-Complex
|
Comment
|
| ## AsTm
|
AsTmE
|
CIV
|
CIV
|
|
| ## Azd
|
CHB
|
|
|
|
- Bioblast links: SUIT protocols - >>>>>>> - Click on [Expand] or [Collapse] - >>>>>>>
- Coupling control
- » Coupling control state
- » ET capacity
- » OXPHOS capacity
- » LEAK respiration
- Pathway control
- » Electron transfer pathway
- » Fatty acid oxidation pathway control state, F
- » NADH electron transfer-pathway state, N
- » Succinate pathway control state, S
- » NS-pathway control state, NS
- » Glycerophosphate pathway control state, Gp
- » Complex IV single step, CIV
- » Anaplerotic pathway control state
- Main fuel substrates
- » Glutamate, G
- » Glycerophosphate, Gp
- » Malate, M
- » Octanoylcarnitine, Oct
- » Pyruvate, P
- » Succinate, S
- Glossary
- » List of SUIT states
- » SUIT concept
Strengths and limitations
- + This protocol provides information on FAO capacity in the absence of other potentially interfering pathways both in LEAK and OXPHOS coupling control states.
- + FNS OXPHOS capacity comprises the most important pathways in many cell types and thus provides a physiologically relevant estimate of maximum mitochondrial respiratory capacity.
- + FNS ET capacity is a good estimate of overal ET capacity in many cell types.
- + This is a good protocol to analyse coupling control at S-pathway, which is an advantage in relation to SUIT-015 and SUIT-017.
- + Glutamate is easier to prepare compared to pyruvate.
- - SRot(E) may be underestimated if S is not saturating.
- - CIV activity is not measured, to save experimental time.
Compare SUIT protocols
MitoPedia concepts:
MiP concept,
SUIT protocol,
Recommended
MitoPedia methods:
Respirometry
Cookies help us deliver our services. By using our services, you agree to our use of cookies.
![]() »Versions
»Versions



