The MitoPedia terminology is developed continuously in the spirit of Gentle Science.
- MitoPedia: SUIT protocols is a component of OROBOROS support.
- See also
- » O2k-Protocols: O2k-Demo experiments
- SUIT: Substrate-uncoupler-inhibitor-titration
| Term | Abbreviation | Description |
|---|---|---|
| Categories of SUIT protocols | SUIT-catg |
Categories of SUIT protocols group SUIT protocols according to all substrate types involved in a protocol (F, N, S, Gp), independent of the sequence of titrations of substrates and inhibitors which define the Electron-transfer-pathway states. The N-type substrates are listed in parentheses, independent of the sequence of titrations. ROX states may or may not be included in a SUIT protocol, which does not change its category. Similarly, the CIV assay may or may not be added at the end of a SUIT protocol, without effect on the category of a SUIT protocol.
|
| Cell respiration | Cell respiration channels metabolic fuels into the chemiosmotic coupling (bioenergetic) machinery of oxidative phosphorylation, being regulated by and regulating oxygen consumption (or consumption of an alternative final electron acceptor) and molecular redox states, ion gradients, mitochondrial (or microbial) membrane potential, the phosphorylation state of the ATP system, and heat dissipation in response to intrinsic and extrinsic energy demands. See also respirometry. In internal or cell respiration in contrast to fermentation, redox balance is maintained by external electron acceptors, transported into the cell from the environment. The chemical potential between electron donors and electron acceptors drives the electron transfer pathway, generating a chemiosmotic potential that in turn drives ATP synthesis. | |
| Coupling-control protocol | CCP | A coupling-control protocol CCP induces different coupling control states at a constant electron-transfer-pathway state. Residual oxygen consumption (Rox) is finally evaluated for Rox correction of flux. The CCP may be extended, when further respiratory states (e.g. cell viability test; CIV assay) are added to the coupling control module consisting of three coupling control states. The term phosphorylation control protocol, PCP, has been introduced synonymous for CCP. » MiPNet article |
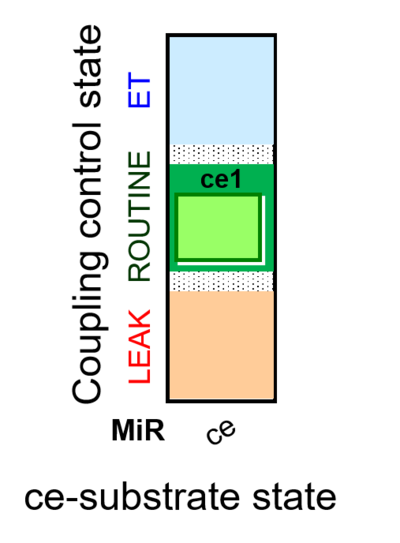
| Coupling-control state | CCS | Coupling-control states are defined in mitochondrial preparations (isolated mitochondria, permeabilized cells, permeabilized tissues, homogenates) as LEAK respiration, OXPHOS, and ET states, with corresponding respiration rates (L, P, E) in any electron-transfer-pathway state which is competent for electron transfer. These coupling states are induced by titration of ADP and uncouplers, and application of specific inhibitors of the phosphorylation pathway. In living cells, the coupling-control states are LEAK respiration, ROUTINE, and ET states of respiration with corresponding rates L, R, E, using membrane-permeable inhibitors of the phosphorylation system (e.g. oligomycin) and uncouplers (e.g. CCCP). Coupling-control protocols induce these coupling-control states sequentially at a constant electron-transfer-pathway state. |
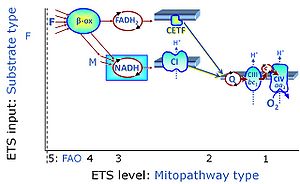
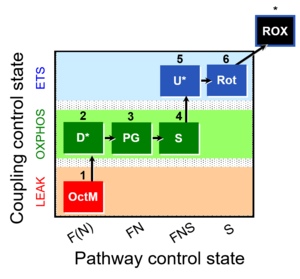
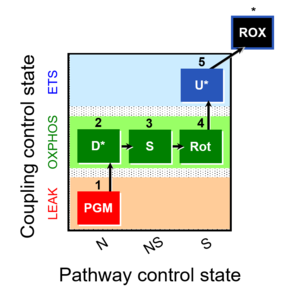
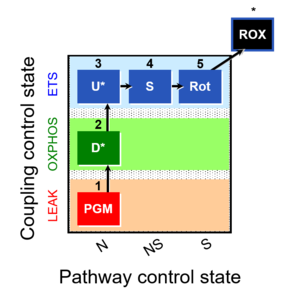
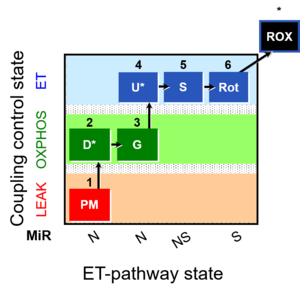
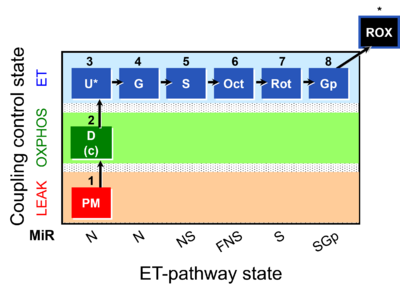
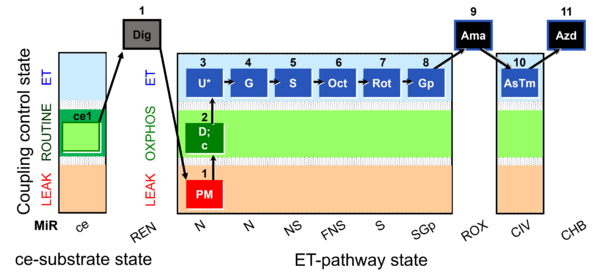
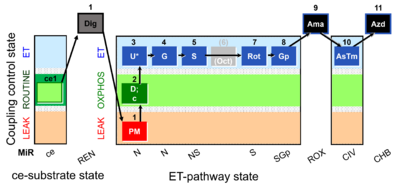
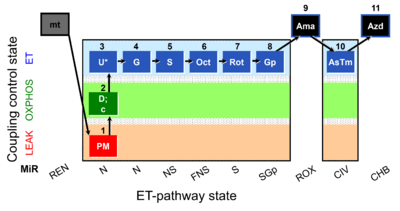
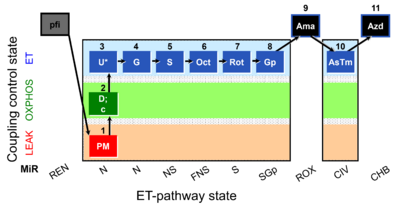
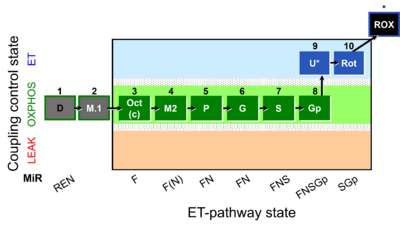
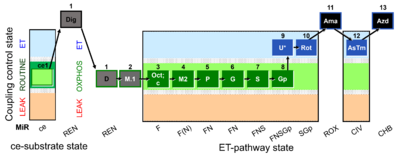
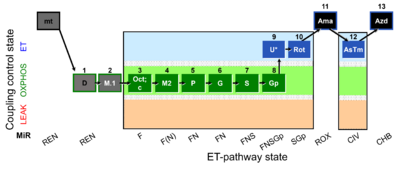
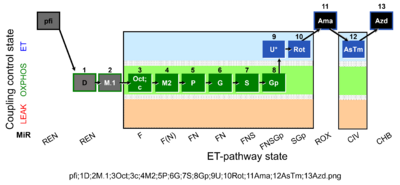
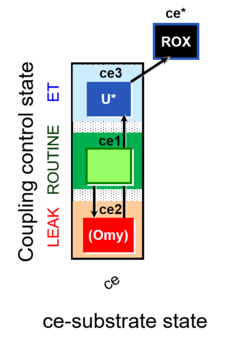
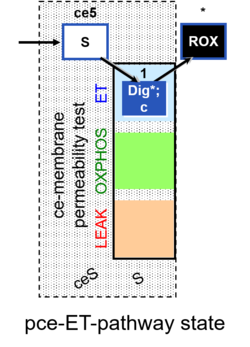
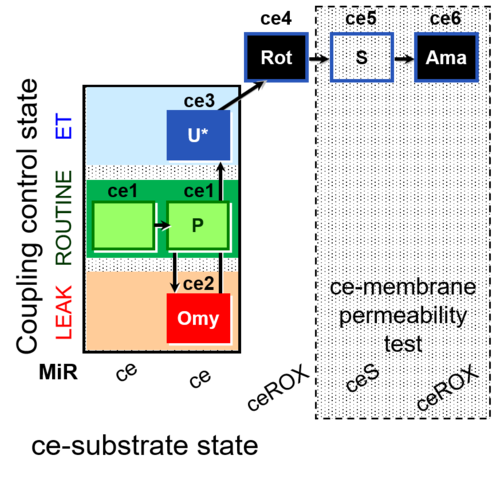
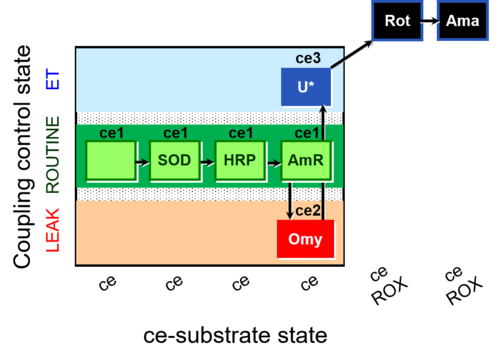
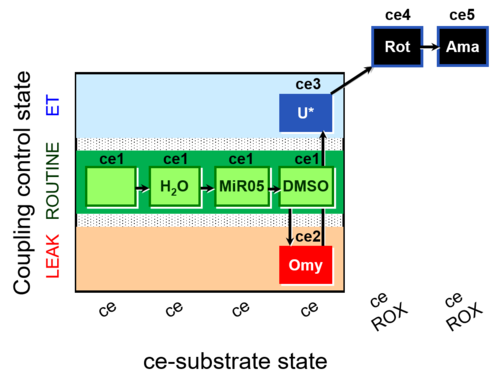
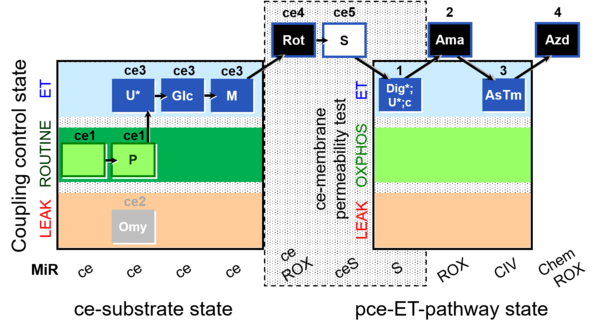
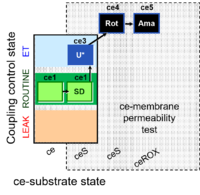
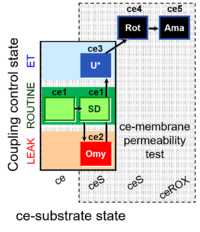
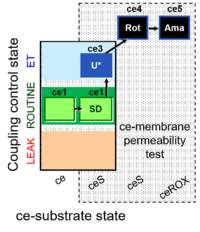
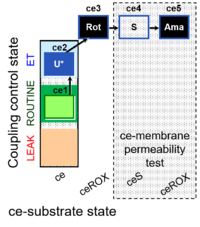
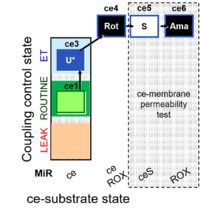
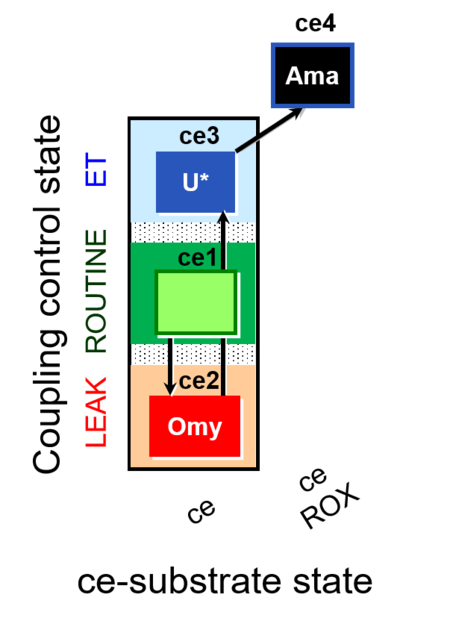
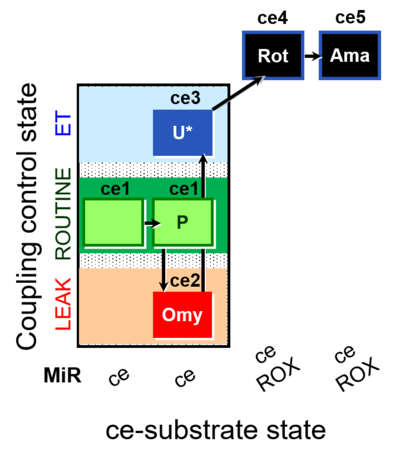
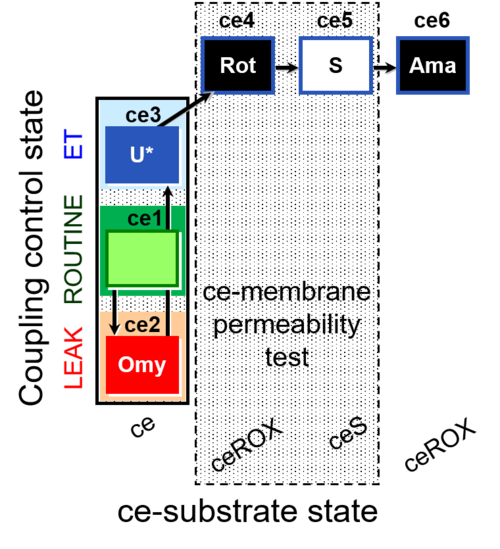
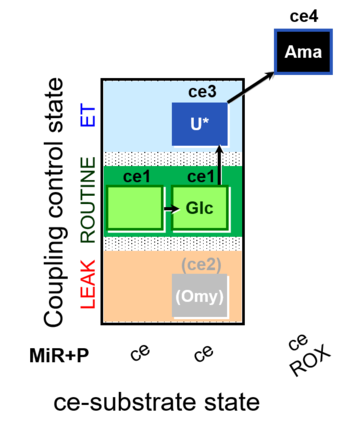
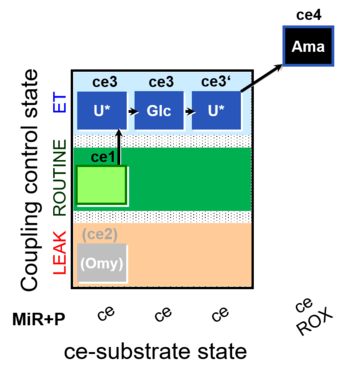
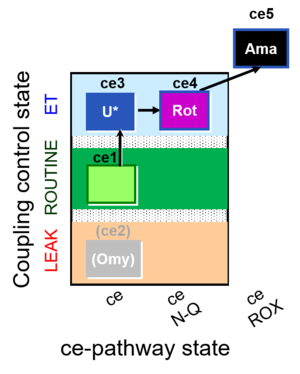
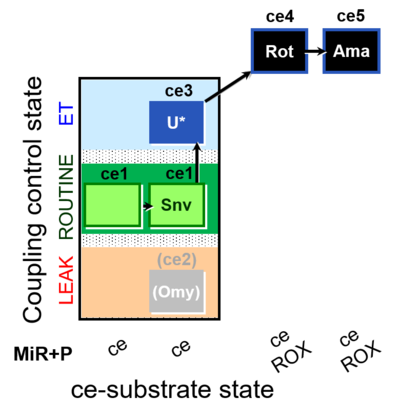
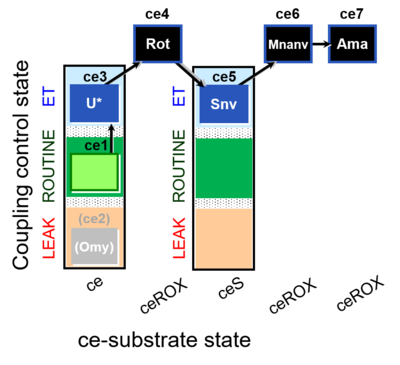
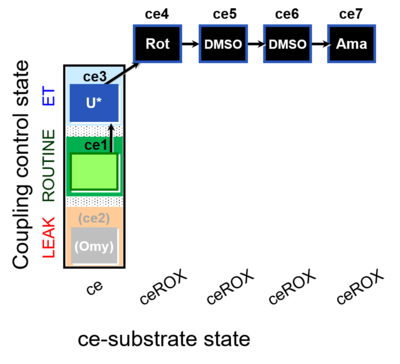
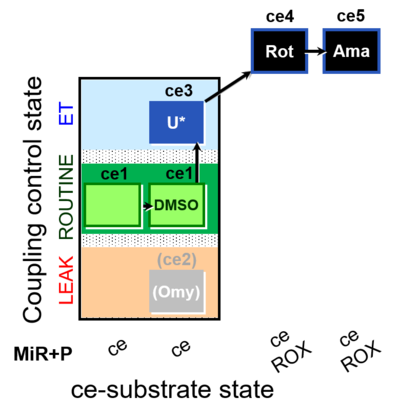
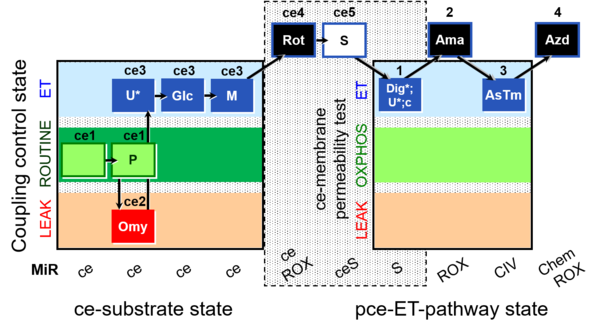
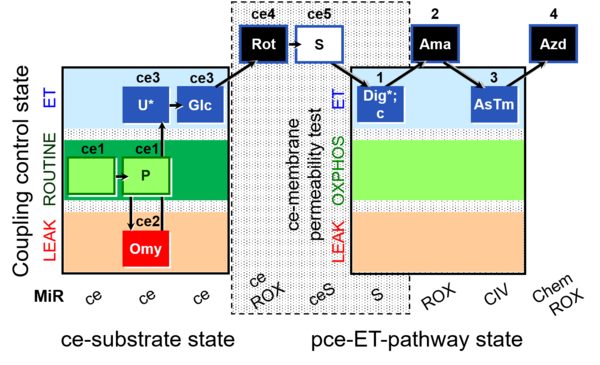
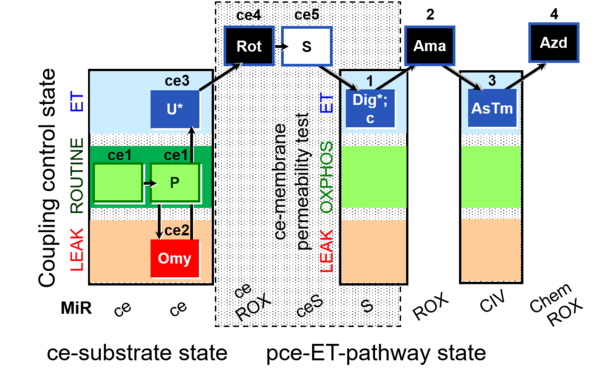
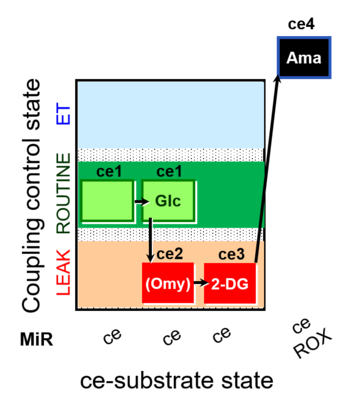
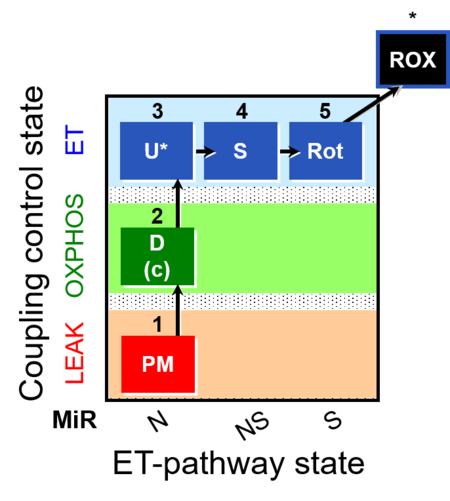
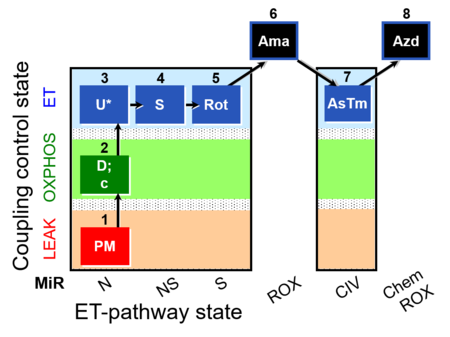
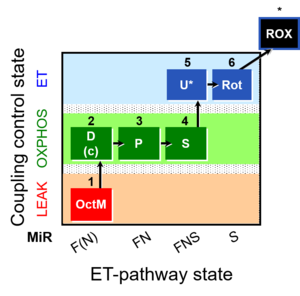
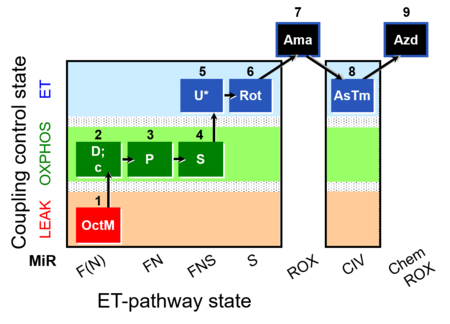
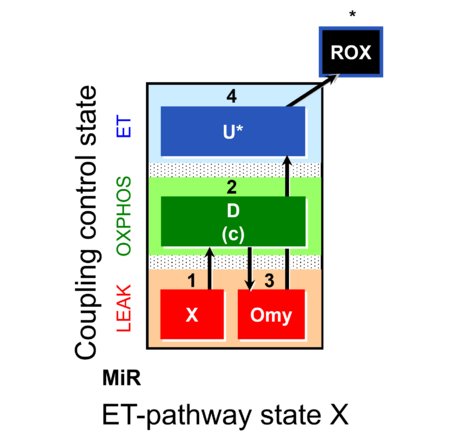
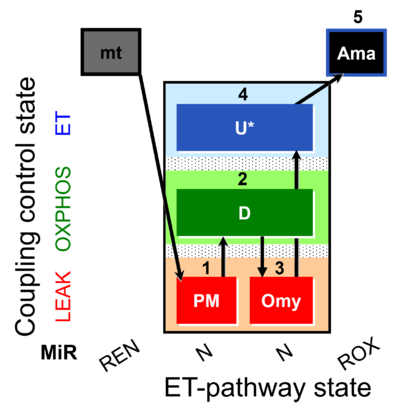
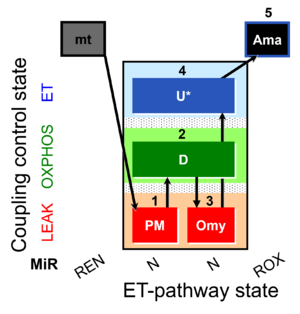
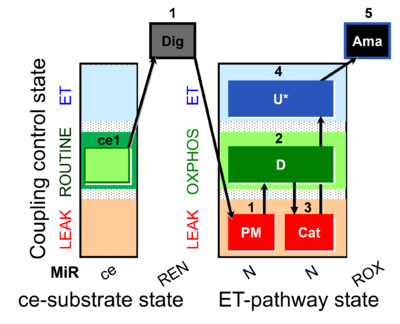
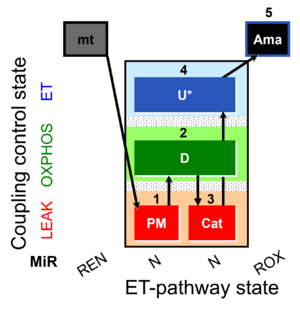
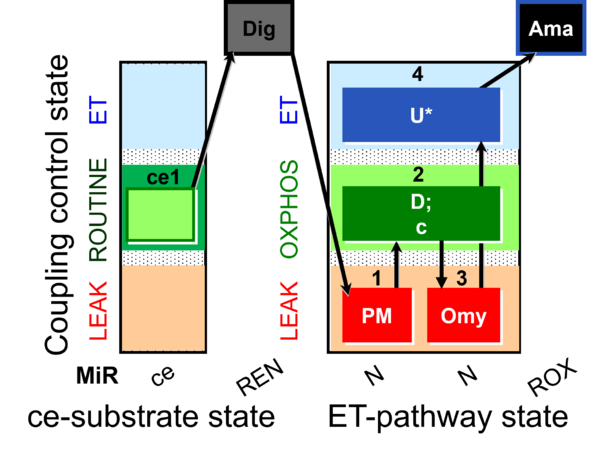
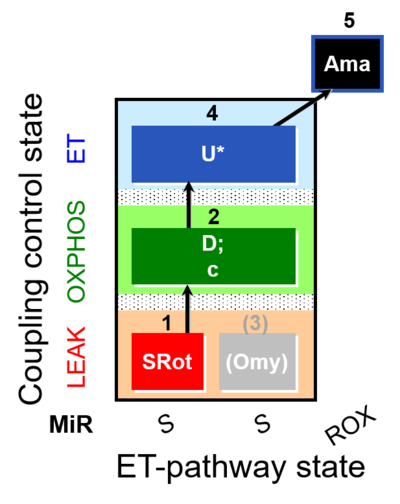
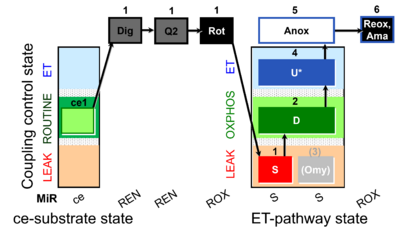
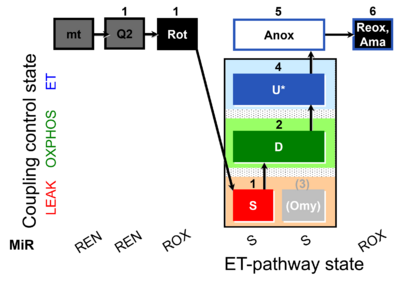
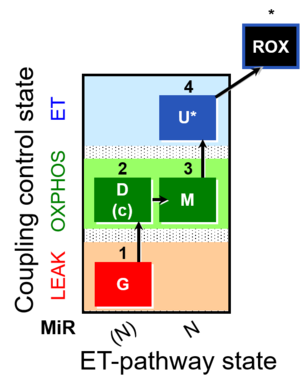
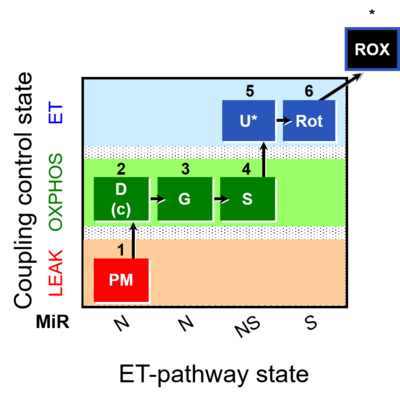
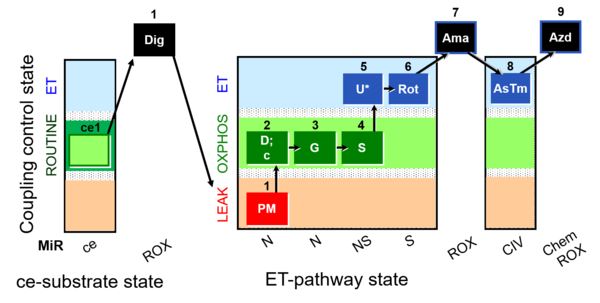
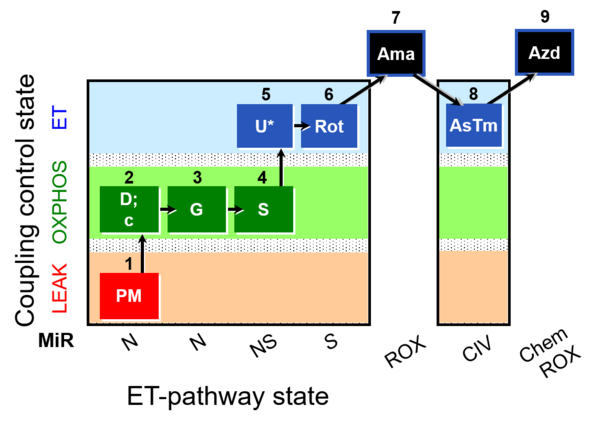
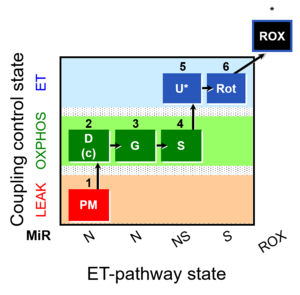
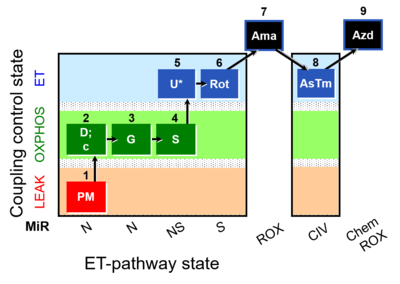
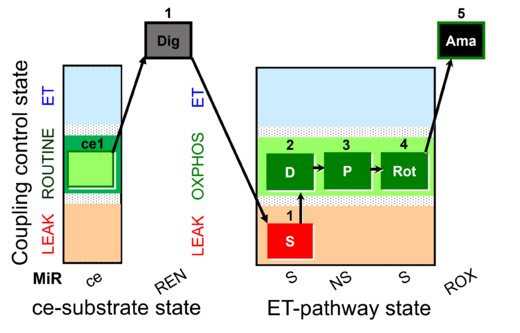
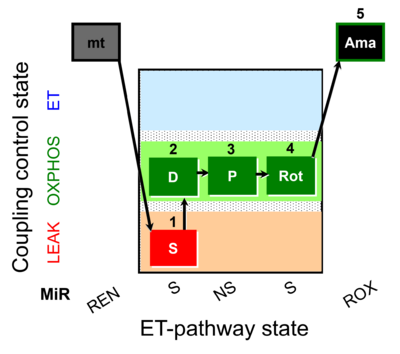
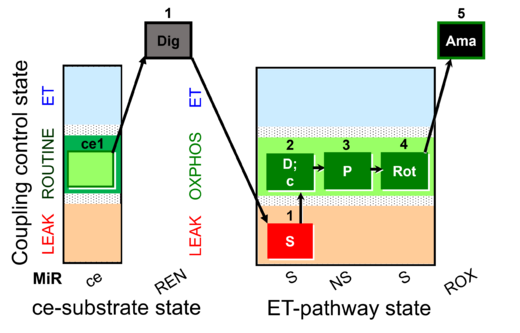
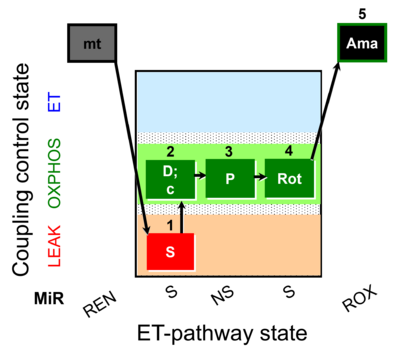
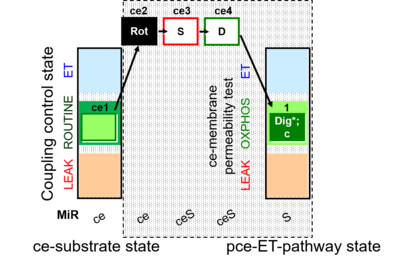
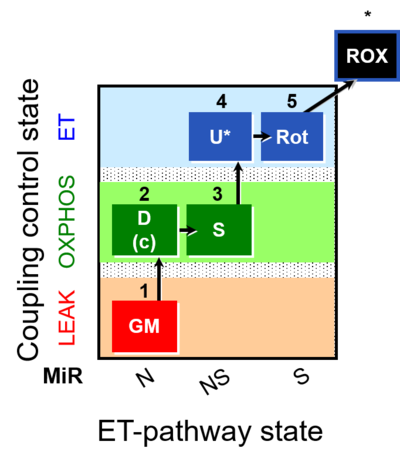
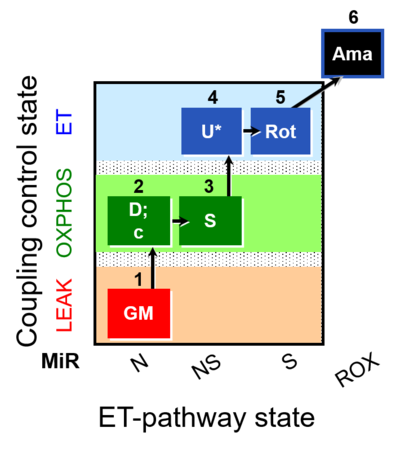
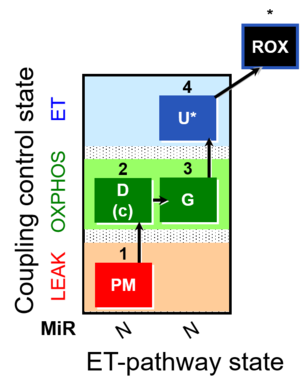
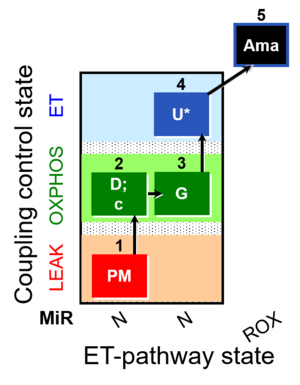
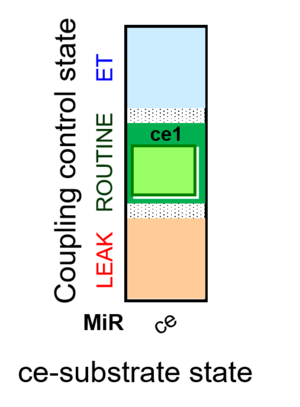
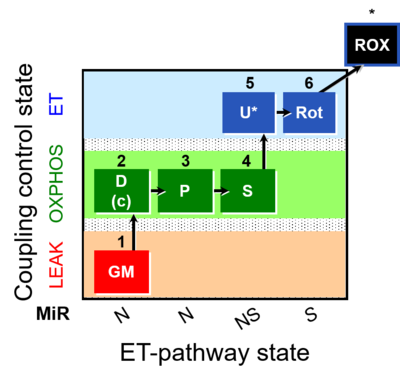
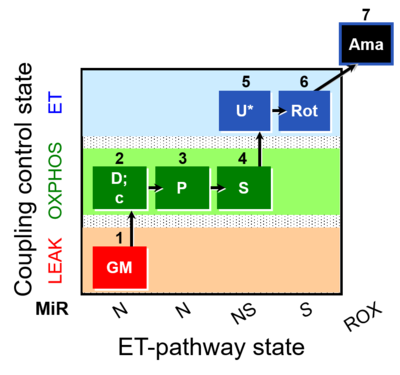
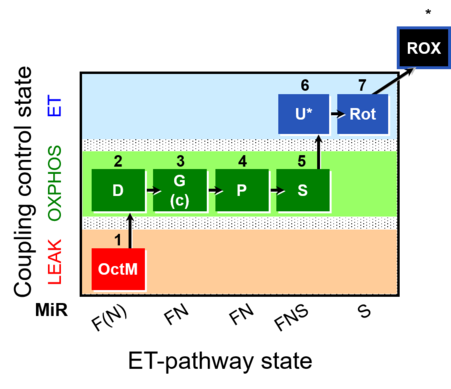
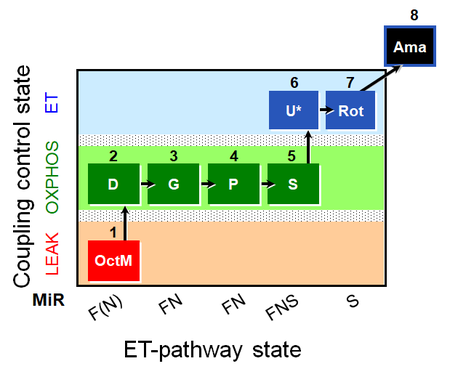
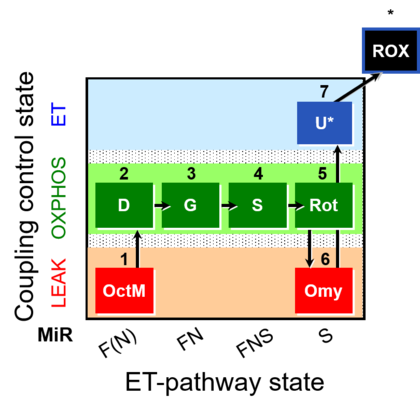
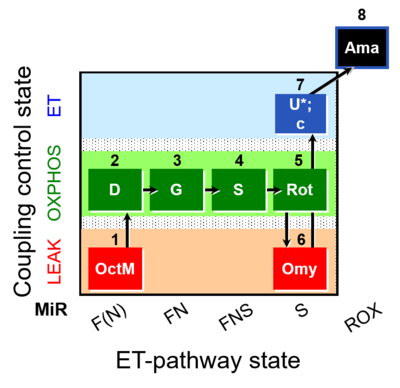
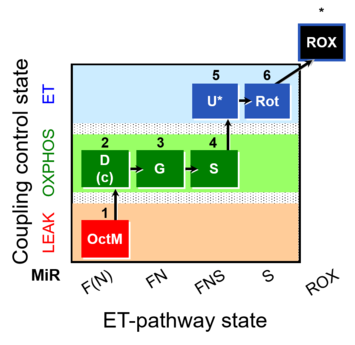
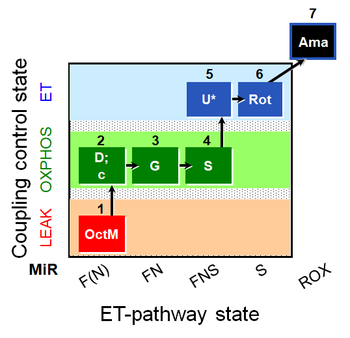
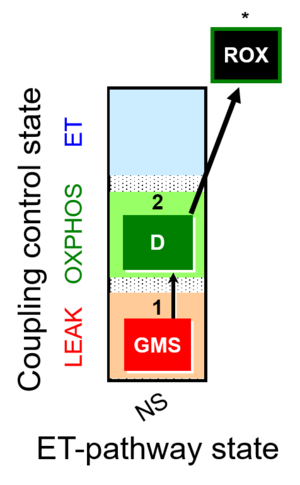
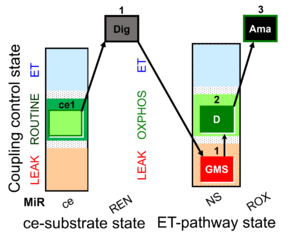
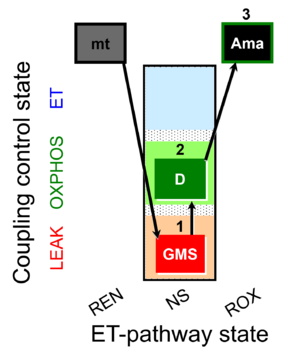
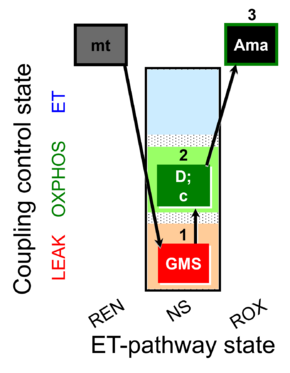
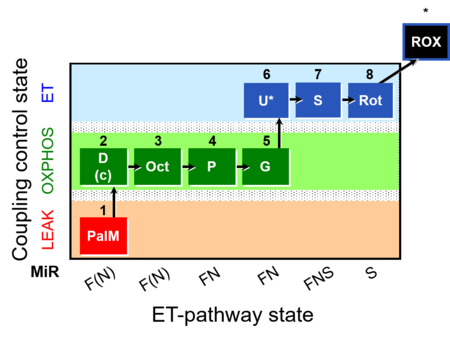
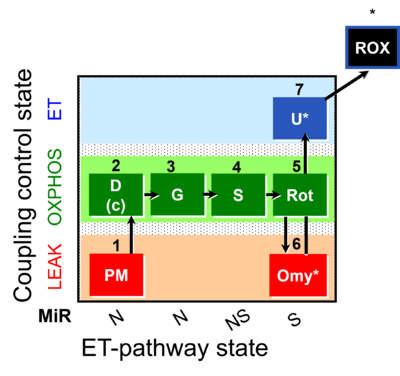
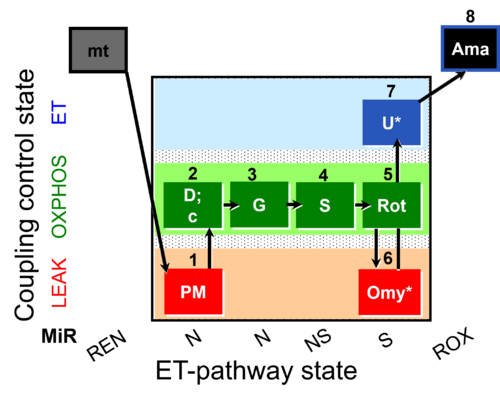
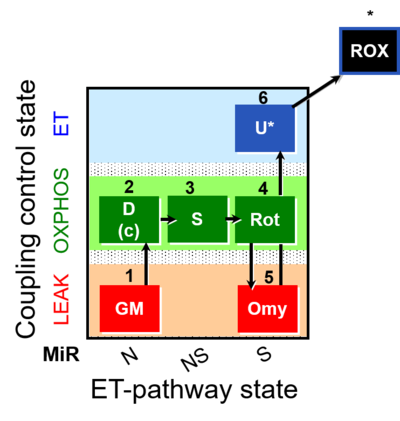
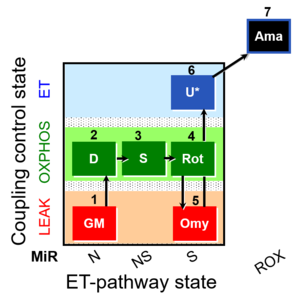
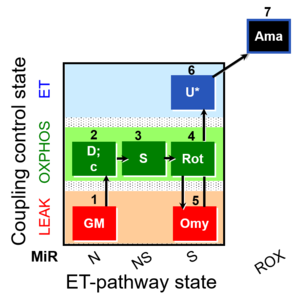
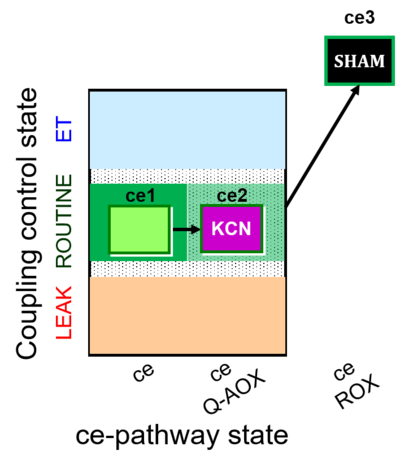
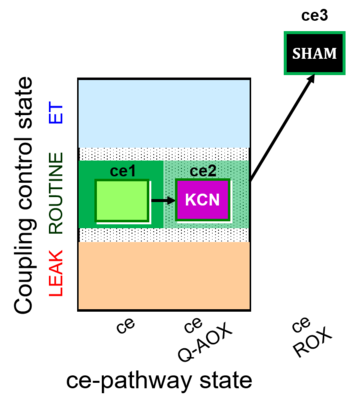
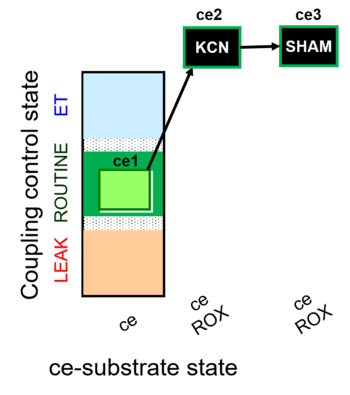
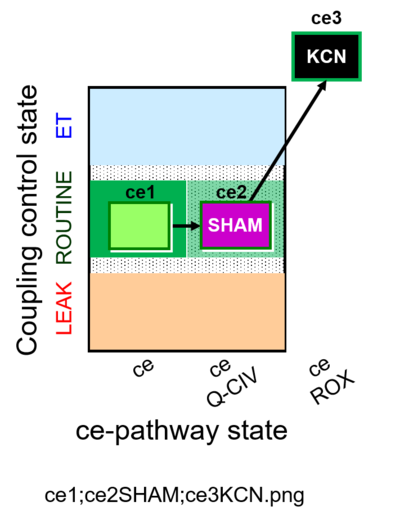
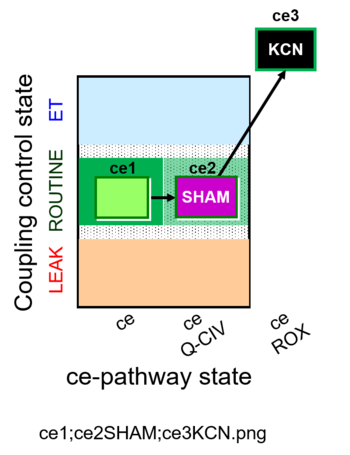
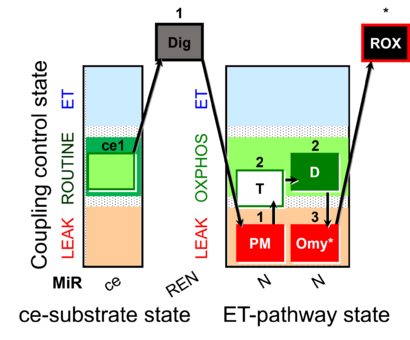
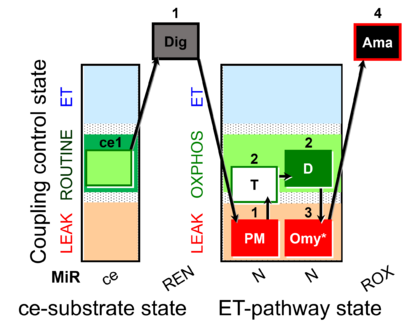
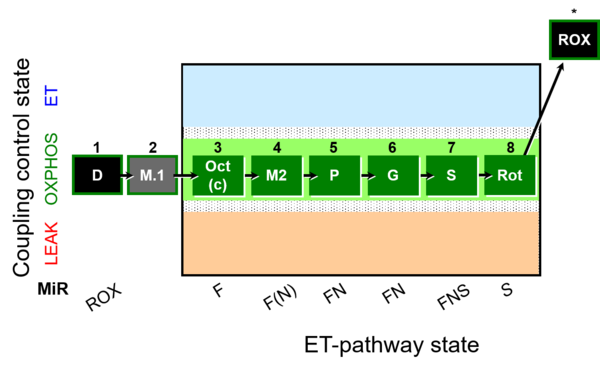
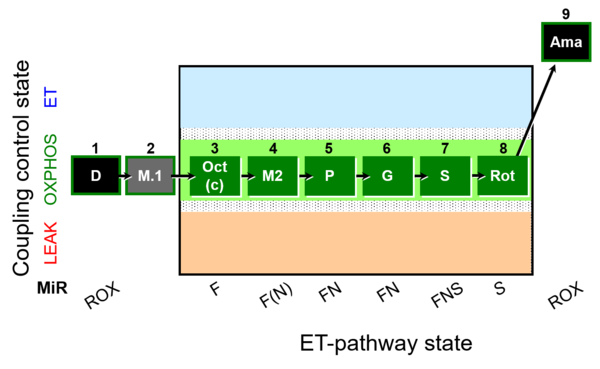
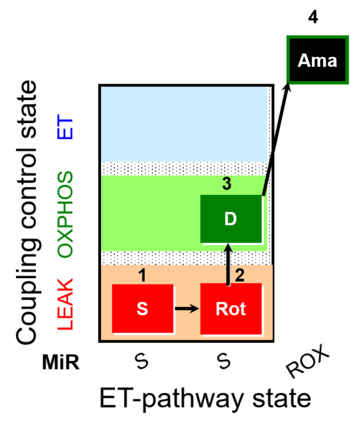
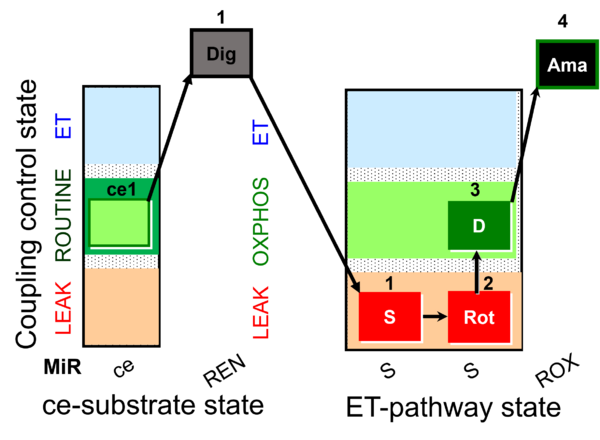
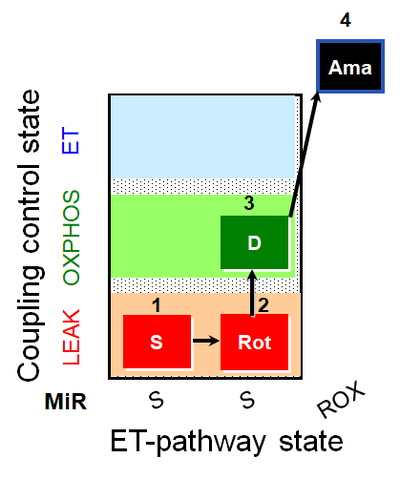
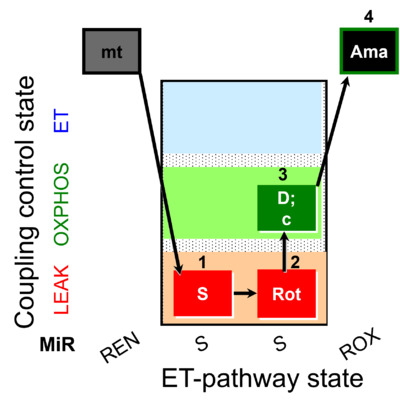
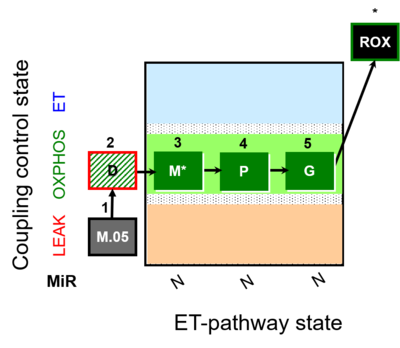
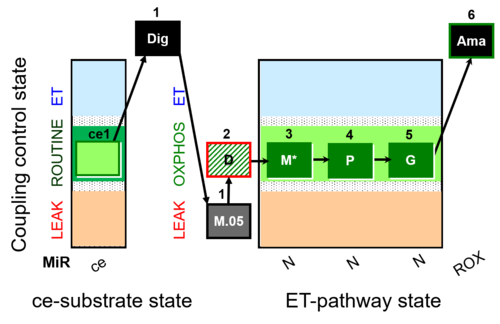
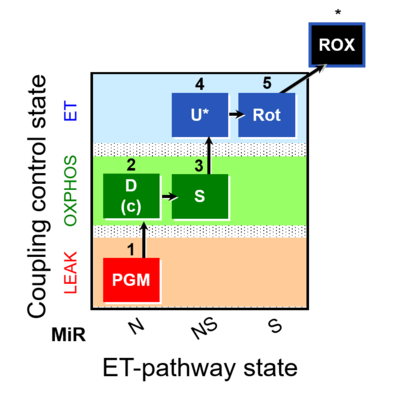
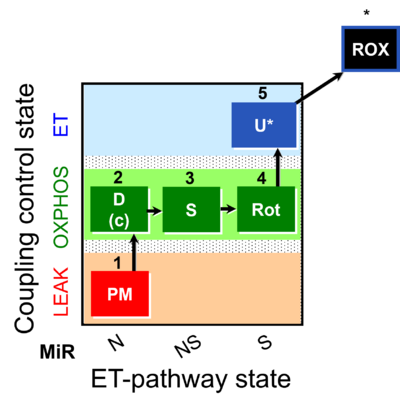
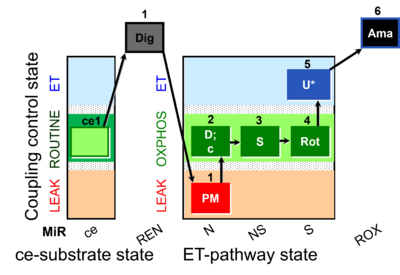
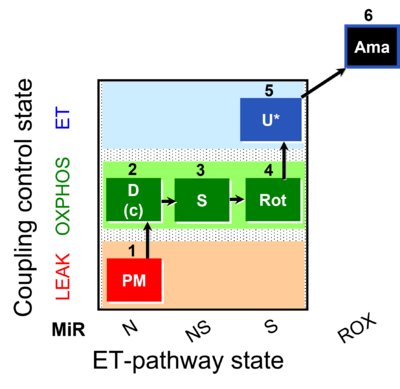
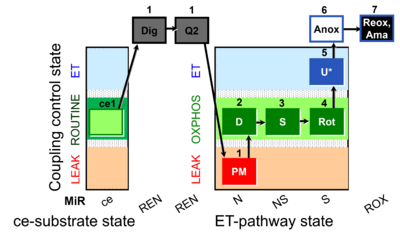
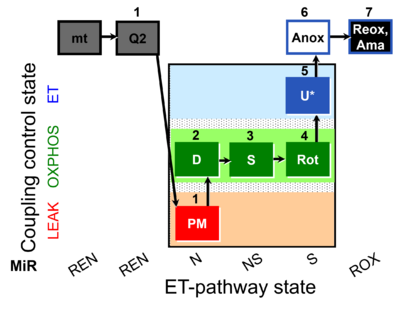
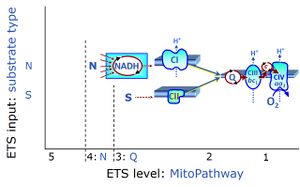
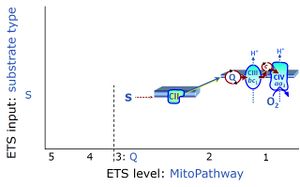
| Coupling/pathway control diagram | CPCD | Coupling/pathway control diagrams illustrate the respiratory states obtained step-by-step in substrate-uncoupler-inhibitor titrations in a SUIT protocol. Each step (to the next state) is defined by an initial state and a metabolic control variable, X. The respiratory states are shown by boxes. X is usually the titrated substance in a SUIT protocol. If X (ADP, uncouplers, or inhibitors of the phosphorylation system, e.g. oligomycin) exerts coupling control, then a transition is induced between two coupling-control states. If X (fuel substrates, e.g. pyruvate and succinate, or Electron transfer pathway inhibitors, e.g. rotenone) exerts pathway control, then a transition is induced between two Electron-transfer-pathway states. The type of metabolic control (X) is shown by arrows linking two respiratory states, with vertical arrows indicating coupling control, and horizontal arrows indicating pathway control. Marks define the section of an experimental trace in a given respiratory state (steady state). Events define the titration of X inducing a transition in the SUIT protocol. The specific sequence of coupling control and pathway control steps defines the SUIT protocol pattern. The coupling/pathway control diagrams define the categories of SUIT protocols (see Figure). |
| Cross-linked respiratory states | CLRS | Coordinated respiratory SUIT protocols are designed to include cross-linked respiratory states, which are common to these protocols. Different SUIT protocols address a variety of respiratory control steps which cannot be accomodated in a single protocol. Cross-linked respiratory states are included in each individual coordinated protocol, such that these states can be considered as replicate measurements, which also allow for harmonization of data obtained with these different protocols. |
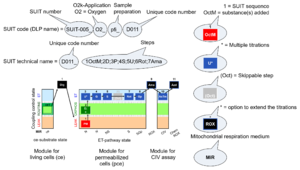
| D-number | D### | D number is the unique code given for each SUIT protocol. In the same SUIT protocol family (SUIT-###), there might be different protocols, specifically designed for different sample type (e.g., different mitochondrial preparations) or for different applications (e.g., O2, AmR, Fluo, MgG). Since the use of different kinds of sample or application may result in slightly different steps, each protocol receives a different D-number. |
| DatLab and SUIT protocols | This is a brief summary of steps to be taken for performing a high-resolution respirometry experiment with SUIT protocols using the OROBOROS Oroboros O2k and DatLab software. (1) Search for a specific SUIT protocol name (go to MitoPedia: SUIT). The list of MitoPedia SUIT protocols can be sorted by categories of SUIT protocols (sorting by SUIT protocol name), which is listed as the 'abbreviation' of the SUIT protocol name. (2) Copy the template for Mark names into your DatLab subdirectory: DatLab\APPDATA\MTEMPLAT. (3) Copy the DatLab-Analysis template for this SUIT protocol. (4) Follow the link to the corresponding publication or MiPNet communication, where the pdf file describing the SUIT protocol is available. (5) A DatLab demo file may be available providing an experimental example. After each sequential titration, a mark is set on the plot for flux or flow. After having set all marks, pull down the 'Mark names' menu, select the corresponding SUIT protocol for mark names, and rename all marks. The Mark names template also provides standard values of the titration volume preceding each mark. (6) Go to 'Mark statistics' [F2], copy to clipboard, and paste into the sample tab in the DatLab-Analysis template.
| |
| ET-pathway substrate types | n.a. | See Electron-transfer-pathway state |
| Electron-transfer-pathway state | ET-pathway state |
Electron-transfer-pathway states are obtained in mitochondrial preparations (isolated mitochondria, permeabilized cells, permeabilized tissues, tissue homogenate) by depletion of endogenous substrates and addition to the mitochondrial respiration medium of fuel substrates (CHNO) activating specific mitochondrial pathways, and possibly inhibitors of specific pathways. Mitochondrial electron-transfer-pathway states have to be defined complementary to mitochondrial coupling-control states. Coupling-control states require ET-pathway competent states, including oxygen supply. Categories of SUIT protocols are defined according to mitochondrial ET-pathway states. » MiPNet article |
| Ethanol | ethanol abs. |
Ethanol or ethyl alcohol, C2H6O or EtOH, is widely used in the laboratory, particularly as a solvent and cleaning agent. There are different grades of high purity ethanol. Up to a purity of 95.6 % ethanol can be separated from water by destillation. Higher concentrations than 95% require usage of additives that disrupt the azeotrope composition and allow further distillation. Ethanol is qualified as "absolute" if it contains no more than one percent water. Whenever 'ethanol abs.' is mentioned without further specification in published protocols, it refers to ≥ 99 % ethanol a.r. (analytical reagent grade).
|
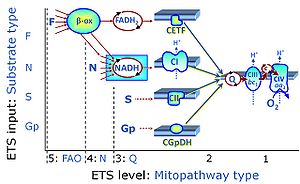
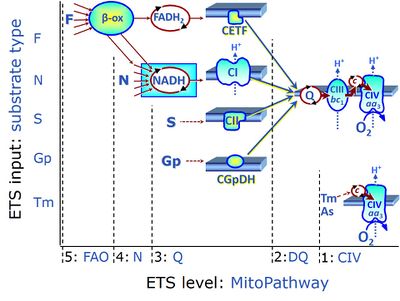
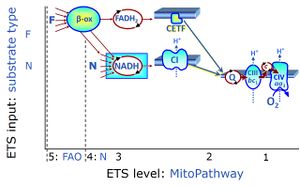
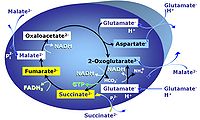
| F-junction | The F-junction is a junction for convergent electron flow in the electron transfer pathway (ET-pathway) from fatty acids through fatty acyl CoA dehydrogenase (reduced form FADH2) to electron transferring flavoprotein (CETF), and further transfer through the Q-junction to Complex III (CIII). The concept of the F-junction and N-junction provides a basis for defining categories of SUIT protocols. Fatty acid oxidation, in the F-pathway control state, not only depends on electron transfer through the F-junction (which is typically rate-limiting) but simultaneously generates NADH and thus depends on N-junction throughput. Hence FAO can be inhibited completely by inhibition of Complex I (CI). In addition and independent of this source of NADH, the N-junction substrate malate is required as a co-substrate for FAO in mt-preparations, since accumulation of AcetylCoA inhibits FAO in the absence of malate. Malate is oxidized in a reaction catalyzed by malate dehydrogenase to oxaloacetate (yielding NADH), which then stimulates the entry of AcetylCoA into the TCA cycle catalyzed by citrate synthase. | |
| Flux control efficiency | jZ-Y | Flux control efficiencies express the control of respiration by a metabolic control variable, X, as a fractional change of flux from YX to ZX, normalized for ZX. ZX is the reference state with high (stimulated or un-inhibited) flux; YX is the background state at low flux, upon which X acts.
Complementary to the concept of flux control ratios and analogous to elasticities of metabolic control analysis, the flux control efficiency of X upon background YX is expressed as the change of flux from YX to ZX normalized for the reference state ZX. » MiPNet article |
| Flux control ratio | FCR | Flux control ratios FCRs are ratios of oxygen flux in different respiratory control states, normalized for maximum flux in a common reference state, to obtain theoretical lower and upper limits of 0.0 and 1.0 (0 % and 100 %). For a given protocol or set of respiratory protocols, flux control ratios provide a fingerprint of coupling and substrate control independent of (1) mt-content in cells or tissues, (2) purification in preparations of isolated mitochondria, and (3) assay conditions for determination of tissue mass or mt-markers external to a respiratory protocol (CS, protein, stereology, etc.). FCR obtained from a single respirometric incubation with sequential titrations (sequential protocol; SUIT protocol) provide an internal normalization, expressing respiratory control independent of mitochondrial content and thus independent of a marker for mitochondrial amount. FCR obtained from separate (parallel) protocols depend on equal distribution of subsamples obtained from a homogenous mt-preparation or determination of a common mitochondrial marker. |
| Harmonized SUIT protocols | H-SUIT | Harmonized SUIT protocols (H-SUIT) are designed to include cross-linked respiratory states. When performing harmonized SUIT protocols in parallel, measurements of cross-linked respiratory states can be statistically evaluated as replicates across protocols. Additional information is obtained on respiratory coupling and substrate control by including respiratory states that are not common (not cross-linked) across the harmonized protocols. |
| Metabolic control variable | X | A metabolic control variable X causes the transition between a background state Y (background rate YX) and a reference state Z (reference rate ZX). X may be a stimulator or activator of flux, inducing the step change from background to reference steady state (Y to Z). Alternatively, X may be an inhibitor of flux, absent in the reference state but present in the background state (step change from Z to Y). |
| MitoFit protocols | MitoFit protocols are moderated by the MitoFit moderators (MitoFit team), either as protocols with direct reference to publications available to the scientific communicty, or protocols additionally described and made available in Bioblast with full information on authors (including contact details), author contributions, and editor (moderator) in charge. This aims at a comprehensive MitoFit data repository, which will require global input and cooperation. | |
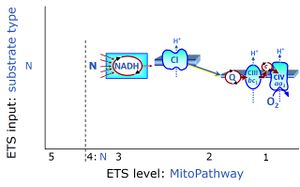
| N-junction | The N-junction is a junction for convergent electron flow in the electron transfer pathway (ET-pathway) from type N substrates (further details »N-pathway control state) through the mt-NADH pool to Complex I (CI), and further transfer through the Q-junction to Complex III (CIII). Representative type N substrates are pyruvate (P), glutamate (G) and malate (M). The corresponding dehydrogenases (PDH, GDH, MDH) and some additional TCA cycle dehydrogenases (isocitrate dehydrogenase, oxoglutarate dehydrogenase generate NADH, the substrate of Complex I (CI). The concept of the N-junction and F-junction provides a basis for defining categories of SUIT protocols based on Electron-transfer-pathway states. | |
| NS e-input | NS, CI&II | NS e-input or the NS-pathway control state is electron input from a combination of substrates for the N-pathway control state and S-pathway control state through Complexes CI and CII simultaneously into the Q-junction. NS e-input corresponds to TCA cycle function in vivo, with convergent electron flow through the Electron transfer pathway. In mt-preparations, NS e-input requires addition not only of NADH- (N-) linked substrates (pyruvate&malate or glutamate&malate), but of succinate (S) simultaneously, since metabolite depletion in the absence of succinate prevents a significant stimulation of S-linked respiration. For more details, see: Additive effect of convergent electron flow. |
| OXPHOS capacity | P | |
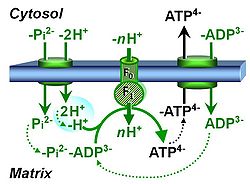
| Phosphorylation pathway | DT | The phosphorylation pathway (phosphorylation system) is the functional unit utilizing the protonmotive force to phosphorylate ADP (D) to ATP (T), and may be defined more specifically as the P»-system. The P»-system consists of adenine nucleotide translocase, phosphate carrier, and ATP synthase. Mitochondrial adenylate kinase, mt-creatine kinase and mt-hexokinase constitute extended components of the P»-system, controlling local AMP and ADP concentrations and forming metabolic channels. Since substrate-level phosphorylation is involved in the TCA-cycle, the P»-system includes succinyl-CoA ligase (GDP to GTP or ADP to ATP). |
| Physiological pathway-control state | See Electron-transfer-pathway state. | |
| Preparation of SUIT chemicals | Preparation of SUIT chemicals describes the preparation of chemicals used in Substrate-Uncoupler-Inhibitor Ttitration (SUIT) protocols. | |
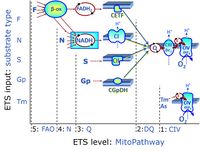
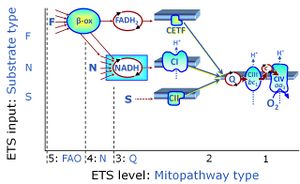
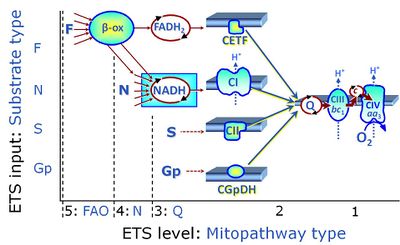
| Q-junction | The Q-junction is a junction for convergent electron flow in the Electron transfer pathway (ET-pathway) from type N substrates and mt-matrix dehydrogenases through Complex I (CI), from type F substrates and FA oxidation through electron-transferring flavoprotein Complex (CETF), from succinate (S) through Complex II (CII), from glycerophosphate (Gp) through glycerophosphate dehydrogenase Complex (CGpDH), from choline through choline dehydrogenase, from dihydro-orotate through dihydro-orotate dehydrogenase, and other enzyme Complexes into the Q-cycle (ubiquinol/ubiquinone), and further downstream to Complex III (CIII) and Complex IV (CIV). The concept of the Q-junction, with the N-junction and F-junction upstream, provides the rationale for defining Electron-transfer-pathway states and categories of SUIT protocols. | |
| Respiratory state | Respiratory states of mitochondrial preparations and living cells are defined in the current literature in many ways and with a diversity of terms. Mitochondrial respiratory states must be defined in terms of both, the coupling-control state and the electron-transfer-pathway state. | |
| SUIT | SUIT | SUIT is the abbreviation for Substrate-Uncoupler-Inhibitor Titration. SUIT protocols are used with mt-preparations to study respiratory control in a sequence of coupling and pathway control states induced by multiple titrations within a single experimental assay. These studies use biological samples economically to gain maximum information with a minimum amount of cells or tissue. |
| SUIT protocol library | SUITs | The Substrate-uncoupler-inhibitor titration (SUIT) protocol library contains a sequential list of SUIT protocols (D001, D002, ..) with links to the specific SUIT pages. Classes of SUIT protocols are explained with coupling and substrate control defined for mitochondrial preparations. |
| SUIT protocol names | SUITp-Names |
The SUIT protocol name starts with (i) the SUIT category which shows the Electron-transfer-pathway states (ET pathway types; e.g. N, S, NS, FNS, FNSGp), independent of the actual sequence of titrations. (ii) A further distinction is provided in the SUIT name by listing in parentheses the substrates applied in the N-pathway control states, again independent of the sequence of titrations, e.g. NS(GM), NS(PM), FNSGp(PGM). (iii) A sequentially selected number is added, e.g. SUIT_FNS(PM)01 (see Coupling/pathway control diagram). The systematic name of a SUIT protocol starts with the SUIT category, followed by an underline dash and the sequence of titration steps (mark names, #X, separated by a comma). The Marks define the section of a respiratory state in the SUIT protocol. The Mark name contains the sequential number and the metabolic control variable, X. The metabolic control variable is the name of the preceding SUIT event. The MitoPedia list of SUIT protocols can be sorted by the short name or the systematic name (hence by SUIT protocol category. The SUIT protocol pattern is best illustrated by a coupling/pathway control diagram. |
| SUIT protocol pattern | SUITp-Pattern | The SUIT protocol pattern describes the type of the sequence of coupling and substrate control steps in a SUIT protocol, which may be liner, orthogonal, or diametral. |
| SUIT reference protocol | SUIT RP | The substrate-uncoupler-inhibitor titration (SUIT) reference protocol, SUIT RP, provides a common baseline for comparison of mitochondrial respiratory control in a large variety of species, tissues and cell types, mt-preparations and laboratories, for establishing a database on comparative mitochondrial phyisology. The SUIT RP consists of two harmonized SUIT protocols (SUIT-001 - RP1 and SUIT-002 - RP2). These are coordinated such that they can be statistically evaluated as replicate measurements of cross-linked respiratory states, while additional information is obtained when the two protocols are conducted in parallel. Therefore, these harmonized SUIT protocols are complementary with their focus on specific respiratory coupling and pathway control aspects, extending previous strategies for respirometrc OXPHOS analysis.
|
| SUITbrowser | Use the SUITbrowser to find the substrate-uncoupler-inhibitor-titration (SUIT) protocol most suitable for addressing your research questions.
Open the SUITbrowser: http://suitbrowser.oroboros.at/
 How to find a DL-Protocol (DLP) How to find a DL-Protocol (DLP) | |
| Substrate control state | See Electron-transfer-pathway state | |
| Substrate-uncoupler-inhibitor titration | SUIT | Mitochondrial Substrate-uncoupler-inhibitor titration (SUIT) protocols are used with mitochondrial preparations to study respiratory control in a sequence of coupling and substrates states induced by multiple titrations within a single experimental assay. |
| Term | Abbreviation | Description |
|---|---|---|
| Anaplerotic pathway control state | a | Anaplerotic pathway control states are fuelled by single substrates which are transported into the mitochondrial matrix and increase the pool of intermediates of the tricarboxylic acid cycle. Malic enzyme (mtME), phosphoenopyruvate carboxykinase (PEPCK), propionyl-CoA carboxylase, and pyruvate carboxylase play important roles in anaplerosis. The glutamate-anaplerotic pathway control state and malate-anaplerotic pathway control state are the most important anaplerotic substrate control states (aN). |
| Complex I&II-linked substrate state | NS | See NS-pathway control state (previous: CI&II-linked) |
| Complex I-linked substrate state | N | See N-pathway control state (previous: CI-linked) versus Complex I |
| Complex II-linked substrate state | SRot, S | See S-pathway control state (previous: CII-linked) |
| Coupling-control state | CCS | Coupling-control states are defined in mitochondrial preparations (isolated mitochondria, permeabilized cells, permeabilized tissues, homogenates) as LEAK respiration, OXPHOS, and ET states, with corresponding respiration rates (L, P, E) in any electron-transfer-pathway state which is competent for electron transfer. These coupling states are induced by titration of ADP and uncouplers, and application of specific inhibitors of the phosphorylation pathway. In living cells, the coupling-control states are LEAK respiration, ROUTINE, and ET states of respiration with corresponding rates L, R, E, using membrane-permeable inhibitors of the phosphorylation system (e.g. oligomycin) and uncouplers (e.g. CCCP). Coupling-control protocols induce these coupling-control states sequentially at a constant electron-transfer-pathway state. |
| Electron-transfer-pathway state | ET-pathway state |
Electron-transfer-pathway states are obtained in mitochondrial preparations (isolated mitochondria, permeabilized cells, permeabilized tissues, tissue homogenate) by depletion of endogenous substrates and addition to the mitochondrial respiration medium of fuel substrates (CHNO) activating specific mitochondrial pathways, and possibly inhibitors of specific pathways. Mitochondrial electron-transfer-pathway states have to be defined complementary to mitochondrial coupling-control states. Coupling-control states require ET-pathway competent states, including oxygen supply. Categories of SUIT protocols are defined according to mitochondrial ET-pathway states. » MiPNet article |
| FN | FN | FN is induced in mt-preparations by addition of NADH-generating substrates (N-pathway control state, or CI-linked pathway control) in combination with one or several fatty acids, which are supplied to feed electrons into the F-junction through fatty acyl CoA dehydrogenase (reduced form FADH2), to electron transferring flavoprotein (CETF), and further through the Q-junction to Complex III (CIII). FAO not only depends on electron transfer through the F-junction (which is typically rate-limiting), but simultaneously generates FADH2 and NADH and thus depends on N-junction throughput. Hence FAO can be inhibited completely by inhibition of Complex I (CI). This physiological substrate combination is required for partial reconstitution of TCA cycle function and convergent electron-input into the Q-junction, to compensate for metabolite depletion into the incubation medium. FS in combination exerts an additive effect of convergent electron flow in most types of mitochondria. |
| FNS | FNS | FNS is induced in mt-preparations by addition of NADH-generating substrates (N-pathway control state, or CI-linked pathway control) in combination with succinate (S-pathway control state; S- or CII-linked) and one or several fatty acids, which are supplied to feed electrons into the F-junction through fatty acyl CoA dehydrogenase (reduced form FADH2), to electron transferring flavoprotein (CETF), and further through the Q-junction to Complex III (CIII). FAO not only depends on electron transfer through the F-junction (which is typically rate-limiting), but simultaneously generates FADH2 and NADH and thus depends on N-junction throughput. Hence FAO can be inhibited completely by inhibition of Complex I (CI). This physiological substrate combination is required for partial reconstitution of TCA cycle function and convergent electron-input into the Q-junction, to compensate for metabolite depletion into the incubation medium. FNS in combination exerts an additive effect of convergent electron flow in most types of mitochondria. |
| FNSGp | FNSGp |
MitoPathway control state: FNSGp
SUIT protocol: SUIT-002 This substrate combination supports convergent electron flow to the Q-junction. |
| Fatty acid oxidation pathway control state | F, FAO | In the fatty acid oxidation pathway control state (F- or FAO-pathway), one or several fatty acids are supplied to feed electrons into the F-junction through fatty acyl CoA dehydrogenase (reduced form FADH2), to electron transferring flavoprotein (CETF), and further through the Q-junction to Complex III (CIII). FAO not only depends on electron transfer through the F-junction (which is typically rate-limiting relative to the N-pathway branch), but simultaneously generates FADH2 and NADH and thus depends on N-junction throughput. Hence FAO can be inhibited completely by inhibition of Complex I (CI). In addition and independent of this source of NADH, the type N substrate malate is required at low concentration (0.1 mM) as a co-substrate for FAO in mt-preparations, since accumulation of Acetyl-CoA inhibits FAO in the absence of malate. Malate is oxidized in a reaction catalyzed by malate dehydrogenase to oxaloacetate (yielding NADH), which then stimulates the entry of Acetyl-CoA into the TCA cycle catalyzed by citrate synthase. Peroxysomal β-oxidation carries out few β-oxidation cycles, thus shortening very-long-chain fatty acids (>C20) for entry into mitochondrial β-oxidation. Oxygen consumption by peroxisomal acyl-CoA oxidase is considered as residual oxygen consumption rather than cell respiration. |
| GM-pathway control state | GM | GM: Glutamate & Malate.
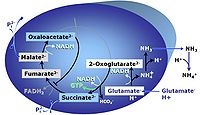
MitoPathway control state: NADH electron transfer-pathway state The GM-pathway control state (glutamate-malate pathway control state) is established when glutamate&malate are added to isolated mitochondria, permeabilized cells and other mitochondrial preparations. Glutamate and transaminase are responsible for the metabolism of oxaloacetate, comparable to the metabolism with acetyl-CoA and citrate synthase. |
| GMS-pathway control state | GMS | GMS: Glutamate & Malate & Succinate.
MitoPathway control: NS Transaminase catalyzes the reaction from oxaloacetate to 2-oxoglutarate, which then establishes a cycle without generation of citrate. OXPHOS is higher with GS (CI&II) compared to GM (CI) or SRot (CII). This documents an additive effect of convergent CI&II electron flow to the Q-junction, with consistent results obtained with permeabilized muscle fibres and isolated mitochondria (Gnaiger 2009). |
| Glutamate-anaplerotic pathway control state | G | G: Glutamate is an anaplerotic NADH-linked type 4 substrate (N). When supplied as the sole fuel substrate in the glutamate-anaplerotic pathway control state, G is transported by the electroneutral glutamate-/OH- exchanger, and is oxidised via mt-glutamate dehydrogenase in the mitochondrial matrix. The G-pathway plays an important role in glutaminolysis. |
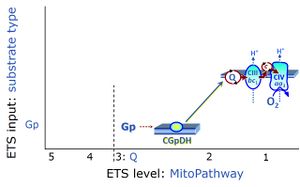
| Glycerophosphate pathway control state | Gp | The glycerophosphate pathway control state (Gp) is an ET-pathway level 3 control state, supported by the fuel substrate glycerophosphate and electron transfer through glycerophosphate dehydrogenase Complex into the Q-junction. The glycerolphosphate shuttle represents an important pathway, particularly in liver and blood cells, of making cytoplasmic NADH available for mitochondrial oxidative phosphorylation. Cytoplasmic NADH reacts with dihydroxyacetone phosphate catalyzed by cytoplasmic glycerophos-phate dehydrogenase. On the outer face of the inner mitochondrial membrane, mitochondrial glycerophosphate dehydrogenase oxidises glycerophosphate back to dihydroxyacetone phosphate, a reaction not generating NADH but reducing a flavin prosthesic group. The reduced flavoprotein donates its reducing equivalents to the electron transfer-pathway at the level of CoQ. |
| LEAK respiration | L | |
| Malate-anaplerotic pathway control state | M | M: Malate alone does not support respiration of mt-preparations if oxaloacetate cannot be metabolized further in the absence of a source of acetyl-CoA. Transport of oxaloacetate across the inner mt-membrane is restricted particularly in liver. Mitochondrial citrate and 2-oxoglutarate (α-ketoglutarate) are depleted by antiport with malate. Succinate is lost from the mitochondria through the dicarboxylate carrier. OXPHOS capacity with malate alone is only 1.3% of that with Pyruvate&Malate in isolated rat skeletal muscle mitochondria. However, many mammalian and non-mammalian mitochondria have a mt-isoform of NADP+- or NAD(P)+-dependent malic enzyme (mtME), the latter being particularly active in proliferating cells. Then the anaplerotic pathway control state with malate alone (aN) supports high respiratory activities comparable to the NADH-linked pathway control states (N) with pyruvate&malate or glutamate&malate substrate combinations (PM-pathway control state, GM-pathway control state). |
| NADH electron transfer-pathway state | N | The NADH electron transfer-pathway state (N) is obtained by addition of NADH-linked substrates (CI-linked), feeding electrons into the N-junction catalyzed by various mt-dehydrogenases. N-supported flux is induced in mt-preparations by the addition of NADH-generating substrate combinations of pyruvate (P), glutamate (G), malate (M), oxaloacetate (Oa), oxoglutarate (Og), citrate, hydroxybutyrate. These N-junction substrates are (indirectly) linked to Complex I by the corresponding dehydrogenase-catalyzed reactions reducing NAD+ to NADH+H+ + H+. The most commonly applied N-junction substrate combinations are: PM, GM, PGM. The malate-anaplerotic pathway control state (M alone) is a special case related to malic enzyme (mtME). The glutamate-anaplerotic pathway control state (G alone) supports respiration through glutamate dehydrogenase (mtGDH). Oxidation of tetrahydrofolate is a NAD(P)H linked pathway with formation of formate. In mt-preparations, succinate dehydrogenase (SDH; CII) is largely substrate-limited in N-linked respiration, due to metabolite depletion into the incubation medium. The residual involvement of S-linked respiration in the N-pathway control state can be further suppressed by the CII-inhibitor malonic acid). In the N-pathway control state ET pathway level 4 is active. |
| NS-pathway control state | NS, CI&II | NS-pathway control is exerted in the NS-linked substrate state (flux in the NS-linked substrate state, NS; or Complex I&II, CI&II-linked substrate state). NS-OXPHOS capacity provides an estimate of physiologically relevant maximum mitochondrial respiratory capacity. NS is induced in mt-preparations by addition of NADH-generating substrates (N-pathway control state in combination with succinate (Succinate pathway; S). Whereas NS expresses substrate control in terms of substrate types (N and S), CI&II defines the same concept in terms of convergent electron transfer to the Q-junction (pathway control). NS is the abbreviation for the combination of NADH-linked substrates (N) and succinate (S). This physiological substrate combination is required for partial reconstitution of TCA cycle function and convergent electron-input into the Q-junction, to compensate for metabolite depletion into the incubation medium. NS in combination exerts an additive effect of convergent electron flow in most types of mitochondria. |
| NSGp | NSGp |
MitoPathway control state: NSGp
SUIT protocol: SUIT-038 This substrate combination supports convergent electron flow to the Q-junction. |
| OXPHOS capacity | P | |
| OctGM | OctGM | OctGM: Octanoylcarnitine & Glutamate & Malate.
MitoPathway control state: FN SUIT protocols: SUIT-015, SUIT-016, SUIT-017 |
| OctGMS | OctGMS | OctGMS: Octanoylcarnitine &Glutamate & Malate& Succinate.
MitoPathway control state: FNS SUIT protocols: SUIT-016, SUIT-017 |
| OctM pathway control state | OctM | OctM: Octanoylcarnitine & Malate.
MitoPathway control state: F SUIT protocols: SUIT-002, SUIT-015, SUIT-016, SUIT-017 Respiratory stimulation of the FAO-pathway, F, by fatty acid FA in the presence of malate M. Malate is a type N substrate (N), required for the F-pathway. In the presence of anaplerotic pathways (e.g., mitochondrial malic enzyme, mtME) the F-pathway capacity is overestimated, if there is an added contribution of NADH-linked respiration, F(N) (see SUIT-002). The FA concentration has to be optimized to saturate the FAO-pathway, without inhibiting or uncoupling respiration. Low concentration of malate, typically 0.1 mM, does not saturate the N-pathway; but saturates the F-pathway. High concentration of malate, typically 2 mM, saturates the N-pathway. |
| OctPGM pathway control state | OctPGM | OctPGM: Octanoylcarnitine & Pyruvate & Glutamate & Malate.
MitoPathway control state: FN SUIT protocols: SUIT-002
|
| OctPGMS pathway control state | OctPGMS | OctPGMS: Octanoylcarnitine & Pyruvate & Glutamate & Malate & Succinate.
MitoPathway control state: FNS SUIT protocol: SUIT-001, SUIT-002, SUIT-015 This substrate combination supports convergent electron flow to the Q-junction. |
| OctPGMSGp pathway control state | OctPGMSGp | OctPGMSGp: Octanoylcarnitine & Pyruvate & Glutamate & Malate & Succinate & Glycerophosphate.
MitoPathway control state: FNSGp SUIT protocol: SUIT-002 This substrate combination supports convergent electron flow to the Q-junction. |
| OctPM pathway control state | OctPM | OctPM: Octanoylcarnitine & Pyruvate & Malate.
MitoPathway control state: FN SUIT protocol: SUIT-002, SUIT-005 This substrate combination supports N-linked flux which is typically higher than FAO capacity (F/FN<0 in the OXPHOS state). In SUIT-RP1, PMOct is induced after PM(E), to evaluate any additive effect of adding Oct. In SUIT-RP2, FAO OXPHOS capacity is measured first, testing for the effect of increasing malate concentration (compare malate-anaplerotic pathway control state, M alone), and pyruvate is added to compare FAO as the background state with FN as the reference state. |
| OctPMS | OctPMS | OctPMS: Octanoylcarnitine & Pyruvate & Malate & Succinate.
MitoPathway control state: FNS SUIT protocol: SUIT-005 |
| PGM-pathway control state | PGM | PGM: Pyruvate & Glutamate & Malate.
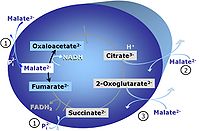
MitoPathway control state: NADH electron transfer-pathway state Pyruvate (P) is oxidatively decarboxylated to acetyl-CoA and CO2, yielding NADH catalyzed by pyruvate dehydrogenase. Malate (M) is oxidized to oxaloacetate by mt-malate dehydrogenase located in the mitochondrial matrix. Condensation of oxaloacate with acetyl-CoA yields citrate (citrate synthase). Glutamate&malate is a substrate combination supporting an N-linked pathway control state, when glutamate is transported into the mt-matrix via the glutamate-aspartate carrier and reacts with oxaloacetate in the transaminase reaction to form aspartate and oxoglutarate. Glutamate as the sole substrate is transported by the electroneutral glutamate-/OH- exchanger, and is oxidized in the mitochondrial matrix by glutamate dehydrogenase to α-ketoglutarate ( 2-oxoglutarate), representing the glutamate-anaplerotic pathway control state. 2-oxoglutarate (α-ketoglutarate) is formed from isocitrate (isocitrate dehydrogenase, from oxaloacetate and glutamate by the transaminase, and from glutamate by the glutamate dehydrogenase. |
| PGMS-pathway control state | PGMS | PGMS: Pyruvate & Glutamate & Malate & Succinate.
MitoPathway control state: NS-pathway control state 2-oxoglutarate is produced through the citric acid cycle from citrate by isocitrate dehydrogenase, from oxaloacetate and glutamate by the transaminase, and from glutamate by the glutamate dehydrogenase. If the 2-oxoglutarate carrier does not outcompete these sources of 2-oxoglutarate, then the TCA cycle operates in full circle with external pyruvate&malate&glutamate&succinate |
| PGMSGp pathway control state | PGMSGp | PGMSGp: Pyruvate & Glutamate & Malate & Succinate & Glycerophosphate.
MitoPathway control state: NSGp SUIT protocol: SUIT-038 This substrate combination supports convergent electron flow to the Q-junction. |
| PM-pathway control state | PM | PM: Pyruvate & Malate.
MitoPathway control state: NADH Electron transfer-pathway state
|
| PMS-pathway control state | PMS | PMS: Pyruvate & Malate & Succinate.
MitoPathway control: CI&II Pyruvate (P) is oxidatively decarboxylated to acetyl-CoA and CO2, yielding NADH catalyzed by pyruvate dehydrogenase. Malate (M) is oxidized to oxaloacetate by mt-malate dehydrogenase located in the mitochondrial matrix. Condensation of oxaloacate with acetyl-CoA yields citrate (citrate synthase). This documents an additive effect of convergent CI&II electron flow to the Q-junction, with consistent results obtained with permeabilized muscle fibres and isolated mitochondria (Gnaiger 2009). |
| PalM | PalM | PalM: Palmitoylcarnitine & Malate.
MitoPathway control state: Fatty acid oxidation pathway control state SUIT protocols: SUIT-019 |
| PalOctM | PalOctM | PalOctM: Palmitoylcarnitine & Octanoylcarnitine & Malate.
MitoPathway control state: Fatty acid oxidation pathway control state SUIT protocols: SUIT-019 |
| PalOctPGM | PalOctPGM | PalOctPGM: Palmitoylcarnitine & Octanoylcarnitine & Pyruvate & Glutamate & Malate.
MitoPathway control state: FN SUIT protocols: SUIT-019 |
| PalOctPGMS | PalOctPGMS | PalOctPGMS: Palmitoylcarnitine & Octanoylcarnitine & Pyruvate & Glutamate & Malate & Succinate.
MitoPathway control state: FNS SUIT protocols: SUIT-019 |
| PalOctPM | PalOctPM | PalOctPM: Palmitoylcarnitine & Octanoylcarnitine & Pyruvate & Malate.
MitoPathway control state: FN SUIT protocols: SUIT-019 |
| PalPGMSGp pathway control state | PalPGMSGp | PalPGMSGp: Palmitoylcarnitine & Pyruvate & Glutamate & Malate & Succinate & Glycerophosphate.
MitoPathway control state: FNSGp SUIT protocol: SUIT-026 This substrate combination supports convergent electron flow to the Q-junction. |
| SGp-pathway control state | SGp | SGp: Succinate & Glycerophosphate.
MitoPathway control state: SGp; obtained with OctPGMSGp(Rot) SUIT protocol: SUIT-001 and ((SUIT-002 |
| Succinate pathway | S, SRot |
The Succinate pathway (S-pathway; S) is the electron transfer pathway that supports succinate-linked respiration (succinate-induced respiratory state; previously used nomenclature: CII-linked respiration; SRot; see Gnaiger 2009 Int J Biochem Cell Biol). The S-pathway describes the electron flux through Complex II (CII; see succinate dehydrogenase, SDH) from succinate and FAD to fumarate and CII-bound flavin adenine dinucleotide (FADH2) to the Q-junction. The S-pathway control state is usually induced in mt-preparations by addition of succinate&rotenone. In this case, only Complex III and Complex IV are involved in pumping protons from the matrix (positive phase, P-phase) to the negative phase (N-phase) with a P»/O2 of 3.5 (P»/O ratio = 1.75). |